Regulatory role of AGC genes in heat stress adaptation in maize (Zea mays)
Abdul Rehman A , Khairiah Mubarak Alwutayd B , Dikhnah Alshehri C , Ibtisam Mohammed Alsudays D , Farrukh Azeem A , Shahroz Rahman A , Muhammad Abid E and Asad Ali Shah
A , Shahroz Rahman A , Muhammad Abid E and Asad Ali Shah  A *
A *
A
B
C
D
E
Abstract
Heat stress represents a significant environmental challenge that restricts maize (Zea mays) growth and yield on a global scale. Within the plant kingdom, the AGC gene family, encoding a group of protein kinases, has emerged as crucial players in various stress responses. Nevertheless, a comprehensive understanding of AGC genes in Z. mays under heat-stress conditions remains elusive. A genome-wide analysis was done using bioinformatics techniques to identify 39 AGC genes in Z. mays, categorising them into three subfamilies based on their conserved domains. We investigated their phylogenetic relationships, gene structures (including intron-exon configurations), and expression patterns. These genes are likely involved in diverse signalling pathways, fulfilling distinct roles when exposed to heat stress conditions. Notably, most ZmAGC1.5, ZmAGC1.9, ZmNDR3, ZmNDR5 and ZmIRE3 exhibited significant changes in expression levels under heat stress, featuring a high G-box ratio. Furthermore, we pinpointed a subset of AGC genes displaying highly coordinated expression, implying their potential involvement in the heat stress response pathway. Our study offers valuable insights into the contribution of AGC genes to Z. mays’s heat stress response, thus facilitating the development of heat-tolerant Z. mays varieties.
Keywords: AGC gene family, AGC subfamilies, bioinformatics techniques, conserved domains, genome-wide analysis, heat stress, protein kinases, stress responses, Zea mays.
Introduction
A family of serine/threonine kinases known as the AGC kinase family is involved in several biological activities, such as cellular division, differentiation, divergence and survival. The name AGC abbreviated for the protein kinases A, G and C, which were the first kinases to be identified as members of this family (Galván-Ampudia and Offringa 2007). The AGC family of kinases is not exclusive to plants; it is found in various organisms across different kingdoms of life. While the term ‘AGC family’ is often associated with plant biology, these kinases are also present in animals, fungi and other eukaryotes (Pearce et al. 2010). The AGC kinases share a conserved catalytic domain but differ in their regulatory domains, which confer specific functions and enable diverse signalling pathways (Leroux et al. 2018). AGCs share a conserved catalytic core with a characteristic bi-lobal structure comprising an N-terminal and a C-terminal lobe. This architecture allows for the recognition and phosphorylation of specific substrates, thereby mediating diverse cellular processes such as cell growth, proliferation and survival. AGC kinases are known for their regulation by multiple phosphorylation sites, including activation loop phosphorylation and hydrophobic motif phosphorylation. Additionally, these kinases often possess regulatory domains that enable integration of various signalling inputs, facilitating precise control of their activity (Pearce et al. 2010). The AGC family is further divided into 10–15 subfamilies based on their regulatory domains and downstream signalling pathways (Bögre et al. 2003). These subfamilies include PDK (phosphoinositide-dependent kinase), AGC1 (protein kinase G), AGC2 (protein kinase C), AGC2-related superfamilies, NDR (nuclear Dbf2-related), IRE (incomplete root hair elongation) and S6K (ribosomal protein S6 kinase) superfamilies (Pearce et al. 2010). Each subfamily has its unique substrate specificity and signalling functions, which are triggered by the diverse range of upstream signalling pathways, also comprising hormones, and second messengers (Zegzouti et al. 2006). In plants, the AGC kinase family and its subfamilies play a diverse part in growth, development and stress responses (Dubé and Tremblay 2005). The AGC1 subfamily regulates plant growth, development and hormone signalling, whereas the AGC2 subfamily is involved in defence against pathogens and pests. AGC3 kinases mediate plant stress responses to abiotic factors. Members of the AGC4 subfamily influence various aspects of plant development, while AGC5 kinases are involved in light signalling and photomorphogenesis (Dhonukshe et al. 2010). The PDK family is involved in the synchronisation of cell persistance and metabolism. PDKs are activated by phosphatidylinositol-3,4,5-trisphosphate (PIP3) and other phosphoinositides and play a key part in activating Akt, a serine/threonine kinase that serves as a key regulator of cell survival and growth (Howe et al. 1992; Anthony et al. 2004; Biondi 2004). The IRE family is a group of AGC kinases that are activated by tyrosine phosphorylation in response to insulin signalling. These kinases play important contribution in insulin signalling and glucose metabolism, and are associated with diseases such as diabetes and cancer (Leroux et al. 2018). The S6K family is involved in the acclimation of protein harmonising and cellular division. Members of this family are actuated by nutritive prompts, and play important part in the regulation of cell growth and proliferation (Mora et al. 2004). The NDR family is involved in the regulation of cell cycle progression and cell morphogenesis. Phosphorylation activates these kinases, and they play a crucial role in regulating cell growth, differentiation, as well as responding to stress and DNA damage. (Tamaskovic et al. 2003). AGC kinases and MAB4/MEL proteins maintain PIN-FORMED proteins (PINs) polarity by limiting lateral diffusion in plant cells. The Arabidopsis thaliana AGC kinases NDR2/4/5 interact with MOB1A/1B and play important roles in pollen development and germination (Zhou et al. 2021). AGC protein kinase AGC1-4 mediates seed size in Arabidopsis (Wang et al. 2020). D6PK plasma membrane polarity requires a repeated CXX(X)P motif and PDK1-dependent phosphorylation (Zhu et al. 2015). Overall, the AGC kinase family and its subfamilies help to regulate important physiological processes in plants. Therefore, understanding the functions and regulatory mechanisms of AGC kinases is important for the further development of novel strategies to advance plant well-being, stress disinclination and crop production (Bögre et al. 2003; Garcia et al. 2012).
Maize (Zea mays), a vital global crop, is a grass plant grown for food, feed and biofuel. Being a C4 plant, it is characterised by excellent productivity in hot and dry conditions due to its distinct photosynthetic pathway (Crafts-Brandner and Salvucci 2002). AGC kinases are known to play a critical role in signalling pathways associated with plant stress responses. Heat stress is an important abiotic stress affecting Z. mays productivity, and understanding the underlying molecular mechanisms is critical for developing heat-tolerant varieties. Previous studies have shown that AGC kinases are involved in a variety of stress responses in plants (Hajibarat and Saidi 2022), suggesting that they may also be involved in heat stress. Morphogenesis-related NDR kinase signalling network is involved in cold tolerance of Z. mays plants (Wu et al. 2022). There are no previous reports on the AGC family kinases in Z. mays. Therefore, This study aims to decipher the specific roles of AGC kinases in Z. mays under heat stress, which may offer insights into molecular processes and potential targets for enhancing heat stress tolerance in Z. mays.
This study examines genome-wide analysis and gene expression of AGC gene family members in Z. mays to understand the regulatory mechanisms contributing to heat tolerance. This insight into highly expressed genes aid in improving Z. mays production under heat stress. The evolutionary relationship categorised the genes and reveal the specific properties of the AGC genes in Z. mays. Understanding heat stress response in context of gene expression is expected to contribute into overall crop improvement (Wilhelm et al. 1999; Harris et al. 2005; Bigini et al. 2021).
Materials and methods
Identification and sequence analysis of AGC gene family in Zea mays
The Arabidopsis Information Resource (TAIR) database (https://www.arabidopsis.org/) (Garnacho-Montero and Amaya-Villar 2010) and the National Center for Biotechnology Information (NCBI) database (https://www.ncbi.nlm.nih.gov/) (Geer et al. 2010) were used to classify members of the AGC gene family. Protein Blast is a widely accepted tool for identifying homologous sequences. The TAIR is specifically designed for Arabidopsis thaliana L., making it a primary resource for researchers studying this model plant and NCBI databases, being comprehensive repositories of genetic information, offer a diverse set of sequences for comparative analysis. Protein blast from NCBI (https://blast.ncbi.nlm.nih.gov) was used (default parameters) to retrieve homologues for the protein sequences of the AGC gene family of Z. mays L.
Phylogenetic analysis of AGC gene family in Z. mays
The phylogenetic tree was built using protein sequences of the AGC gene family from A. thaliana and homologues from Z. mays. The choice of A. thaliana as a reference for Z. mays AGC genes holds significance due to several reasons. A. thaliana is a well-studied model plant with a fully sequenced genome, and its AGC gene repertoire is relatively well-characterised. Using A. thaliana as a reference allows for comparative analysis, leveraging the existing knowledge base to infer potential functions and regulatory mechanisms of Z. mays AGC genes. Furthermore, the evolutionary conservation of AGC genes between A. thaliana and Z. mays provides insights into the functional and structural aspects of these genes. The maximum likelihood tree was generated through default parameters in IQ-TREE (http://iqtree.cibiv.univie.ac.at/) (Nguyen et al. 2015). IQ-TREE, a prominent tool for phylogenetic tree reconstruction, excels in maximum likelihood inference, robustly estimating evolutionary trees for accurate reconstructions. Its advanced model selection and provision of reliable branch support values, alongside a user-friendly interface, make it accessible and efficient for researchers in phylogenetic analyses. In addition, the tree was modified with iTOL (https://itol.embl.de/) (Letunic and Bork 2019).
Gene structures and motif analysis of AGC gene family in Z. mays
The gene strctures of AGC gene family in Z. mays was identified using the online server called Gene Structure Display Server (GSDS, http://gsds.gao-lab.org), which is a bioinformatics tool utilised for visualising and analysing the gene structure of DNA sequences, aiding in the identification of exon–intron boundaries and other structural features. Motifs and their locations were identified using the protein sequences of ZmAGC members. MEME (Multiple Em for Motif Elicitation) is a bioinformatics tool designed for discovering conserved motifs in DNA, RNA or protein sequences. It employs statistical algorithms to identify significant motifs, providing insights into potential functional elements across sequences. Therefore, MEME was employed to perform motif analysis using default settings (https://meme-suite.org/meme/tools/meme) (Bailey et al. 2015).
Physiochemical properties of AGC gene family in Z. mays
ProtParam (https://web.expasy.org/protparam/) was used to predict the physiochemical properties of AGC proteins, such as molecular weight, theoretical pI, number of amino acids and aliphatic index (Garg et al. 2016). Softberry ProtComp tool was used to predict the proteins’ sub-cellular localisation (http://www.softberry.com/berry.phtml?topic=protcomppl&group=programs&subgroup=proloc).
Domain analysis and protein-protein interaction analysis
The identification of conserved domains was carried out using the NCBI database CDD (Conserved Domain Database) available at https://www.ncbi.nlm.nih.gov/Structure/cdd/cdd.shtml, and the Pfam database (Marchler-Bauer et al. 2015). The results obtained from the analysis were visualised using a tool called Tbtools. The protein–protein interaction network (PPI) was built with the help of a STRING database (https://string-db.org) (Mering et al. 2003). The entered sequence data was transferred to this database and a protein interaction with other proteins in the family was built. In addition, the proteins were also divided into three groups using k-means clustering algorithm.
Promoter region and regulatory elements analysis of ZmAGC genes
The promoter and untranslated region (UTR) sections of every single gene in the AGC gene family were gathered and saved in a file from the NCBI nucleotide database. Cis-regulatory elements were subsequently predicted using the PlantCare tool, an online server-based tool that employs the utilising the gene’s promoter and UTR sequences for this analysis (https://bioinformatics.psb.ugent.be/webtools/plantcare/html/) (Lescot et al. 2002). Because PlantCare can only handle only one sequence at a given moment, the process took around an hour or even longer to accomplish for all the genes. Subsequently, the outcomes were aggregated and visualised by utilising TBtools, which is an open-source software available on GitHub (https://github.com/CJ-Chen/TBtools) (Chen et al. 2020).
Gene expression analysis of ZmAGC members under heat stress
Data on heat stress transcriptomes were retrieved from the NCBI SRA database, accession number SRX19031355 (https://www.ncbi.nlm.nih.gov/sra) (Benson et al. 2010). The transcriptome data was then used to map the reads against the Z. mays reference genome Zm-B73-REFERENCE-NAM-5.0 using the Galaxy server. The data retrieved and organised by the RNA-seq tool Galaxy was used, and gene expression levels were assessed using the fragment mapping tool Cufflinks (which specialises in transcription, estimation, and testing of DEGs in RNA samples). This data was then used for expression profiling using another tool called TBtools.
RNA extraction, cDNA synthesis and real-time qPCR of AGC gene family in Z. mays
RNA isolation from Z. mays leaves was conducted using the Trizol method, and quantification was performed with the Thermo Nanodrop 2000. Subsequently, for the 1 mg of the First-Strand Synthesis (FSS) kit was used to synthesise cDNA from RNA. The resulting cDNA was stored at −20°C for future experiments. For quantitative real-time polymerase chain reaction (qPCR), the iTaq Universal SYBR Green Super Mix (Bio-Rad Labs, Hercules, CA, USA) was employed in conjunction with a qPCR (detection system (CFX96 Touch-TM RT PCR detection system). Gene-specific primers were designed using the Oligo Calculator (http://mcb.berkeley.edu/labs/krantz/tools/oligocalc.html/) (Kibbe 2007) and the NCBI Primer BLAST algorithm (https://www.ncbi.nlm.nih.gov/tools/primer-blast/) was used to validate gene-specific primers (Ye et al. 2012). T-test was employed for statistical analysis of expression data.
Targeting of PIN auxin efflux carriers by AGC kinases
The PPI network was assembled with data from the STRING database (https://string-db.org/), and the proteins were subsequently grouped into three distinct clusters using the k-means clustering technique. The protein structures were then predicted using an online server known as iTASSER (https://zhanggroup.org/I-TASSER/). To analyse and compare the protein sequences, we employed the MEGA alignment software (ver. 7, https://megasoftware.net/). In this analysis, the consensus regions or motifs were identified and prominently marked using black boxes.
Results
Identification and sequence analysis of AGC gene family in Z. mays
The process of identifying biological sequences typically encompasses four stages. Initially, a reference sequence is identified, which in our study, involved selecting a member of the AGC Gene family from TAIR. This reference sequence is subsequently employed as a query for conducting a BLAST search against the plant sequence of interest, which, in our case, was Z. mays. The obtained results are then scrutinised for reproducibility and validated by examining a specific signature domain within the plant-specific protein sequence.
A total of 39 reference protein sequences from A. thaliana were retrieved from the TAIR database. Using these sequences, homologues in Z. mays were identified and retrieved from NCBI, resulting in the recognition of 39 homologues for further analysis. All sequences underwent a thorough re-examination to ensure that the conserved Pkinase domain is present. This domain was also discussed in the context of the analysis of AGC protein kinases within this cereal crop, i.e. ZmAGC members.
Phylogenetic analysis of the AGC gene family in Z. mays
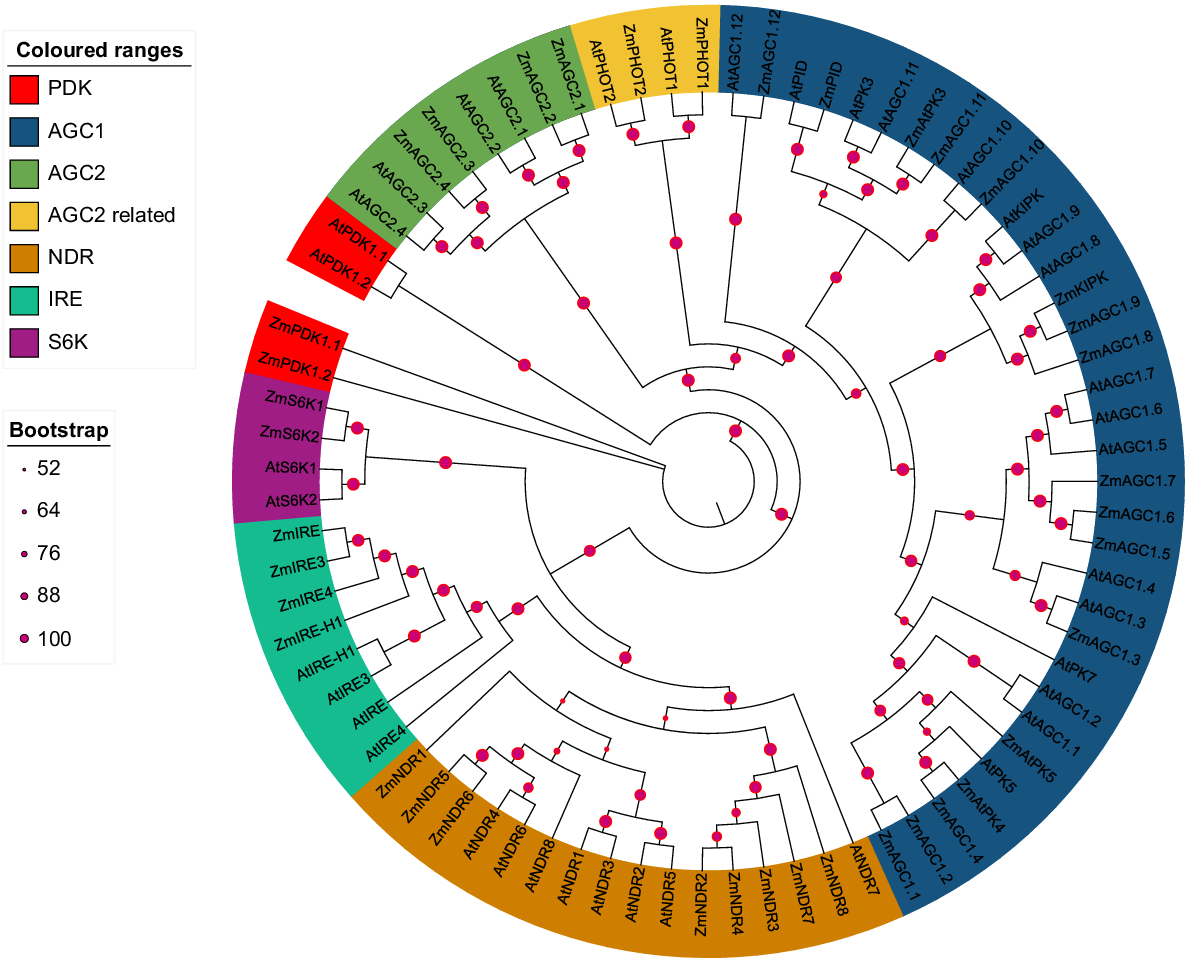
Phylogenetic analysis is a powerful tool for studying the evolutionary relationships of genes or proteins between different species. Using the online IQ-TREE tool, a maximum likelihood parsimony tree was constructed and seven AGC subfamilies were generated. A total of 78 protein sequences were used in the phylogenetic analysis. Phylogenetic analysis of the AGC gene family in Z. mays was performed using Z. mays (ZmAGC) and A. thaliana (AtAGC) proteins. AGC gene sequences from Z. mays were collected and aligned to begin the phylogenetic analysis of AGC genes in Z. mays. Similarly, protein sequences identified in various groups all fall into the same category. In our study, we predicted the existence of several subfamilies within the AGC gene family in Z. mays. Specifically, we identified and counted the following subfamilies and their members: PDK (four members); AGC1 (34 members); AGC2 (eight members); AGC2-associated (four members); NDR (16 members); IRE (eight members); and S6K (four members). By comparing the subfamilies of Z. mays with A. thaliana, we observed a significant expansion in Z. mays. The expansion was most pronounced in subfamilies such as PDK, AGC1, AGC2, and the related AGC2, NDR, IRE and S6K, where the number of members in Z. mays was significantly greater than in A. thaliana. These differences in subfamily size between Z. mays and A. thaliana suggest different evolutionary trajectories or species-specific adaptations within the AGC gene family (Fig. 1).
Physiochemical properties of AGC gene family in Z. mays
Table 1 shows the physiochemical properties of 39 AGC proteins. Our analysis found proteins with higher amino acid numbers and molecular weights. Specifically, ZmIRE-H1, ZmIRE3 and ZmIRE were among the proteins that exhibited these higher values. Theoretical predictions of the isoelectric points (pI) of the proteins gave an average value of 7.62. Furthermore, the proteins were predicted to be located in the nucleus, cytoplasm, plasma membrane and extracellular spaces of the cell, indicating their likely association with activities in the nucleus, cytoplasm, plasma membran, and extracellular spaces. Table 1 contains information on various other properties of the proteins.
| Sr. no. | Protein name | Locus tag | No. of amino acids | Molecular weight | Theoretical pI | Exon | Chromosome no. | GRAVY | Subcellular localisation | |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | ZmPDK1.1 | LOC103650947 | 504 | 55,986.62 | 6.78 | 11 | 3 | −0.38 | Cytoplasm | |
| 2 | ZmPDK1.2 | LOC104567329 | 495 | 55,138.77 | 8.04 | 11 | 3 | −0.37 | Plasma membrane | |
| 3 | ZmS6K1 | LOC100273759 | 481 | 53,432.91 | 6.45 | 7 | 1 | −0.34 | Plasma membrane | |
| 4 | ZmS6K2 | LOC100281573 | 481 | 53,664.29 | 7.7 | 8 | 9 | −0.33 | Plasma membrane | |
| 5 | ZmIRE | LOC103643892 | 1258 | 138,314.46 | 6.09 | 18 | 1 | −0.52 | Nuclear | |
| 6 | ZmIRE-H1 | LOC103625948 | 1261 | 139,109.45 | 6.04 | 19 | 5 | −0.52 | Nuclear | |
| 7 | ZmIRE3 | LOC104578832 | 1260 | 138,541.73 | 6.09 | 18 | 1 | −0.52 | Nuclear | |
| 8 | ZmIRE4 | LOC104784721 | 1119 | 124,494.28 | 5.53 | 18 | 1 | −0.49 | Nuclear | |
| 9 | ZmNDR1 | LOC100381537 | 573 | 65,369.17 | 6.01 | 14 | 1 | −0.69 | Extracellular | |
| 10 | ZmNDR2 | LOC100382815 | 555 | 64,269.4 | 8.84 | 14 | 8 | −0.79 | Nuclear | |
| 11 | ZmNDR3 | LOC100491724 | 556 | 64,340.48 | 8.84 | 14 | 8 | −0.79 | Nuclear | |
| 12 | ZmNDR4 | LOC100285373 | 547 | 62,296.13 | 5.85 | 13 | 8 | −0.52 | Extracellular | |
| 13 | ZmNDR5 | LOC100479234 | 555 | 64,269.4 | 8.84 | 14 | 8 | −0.79 | Nuclear | |
| 14 | ZmNDR6 | LOC100385472 | 547 | 62,230.03 | 5.85 | 13 | 8 | −0.51 | Extracellular | |
| 15 | ZmNDR7 | LOC100483817 | 556 | 64,310.4 | 8.84 | 14 | 8 | −0.79 | Nuclear | |
| 16 | ZmNDR8 | LOC100193886 | 556 | 64,392.77 | 8.97 | 13 | 3 | −0.77 | Nuclear | |
| 17 | ZmAGC1.1 | LOC542271 | 572 | 62,058.69 | 8.04 | 3 | 2 | −0.29 | Cytoplasm | |
| 18 | ZmAGC1.2 | LOC542372 | 572 | 62,048.65 | 8.04 | 3 | 2 | −0.29 | Cytoplasm | |
| 19 | ZmAGC1.3 | LOC100285441 | 685 | 74,622.5 | 6.15 | 5 | 10 | −0.31 | Plasma membrane | |
| 20 | ZmAGC1.4 | LOC100192540 | 577 | 63,207.38 | 7.54 | 9 | 2 | −0.51 | Plasma membrane | |
| 21 | ZmAGC1.5 | LOC103635710 | 530 | 57,619.51 | 6.79 | 2 | 8 | −0.42 | Cytoplasm | |
| 22 | ZmAGC1.6 | LOC104675812 | 446 | 48,773.39 | 6.9 | 7 | 8 | −0.2 | Cytoplasm | |
| 23 | ZmAGC1.7 | LOC103649538 | 532 | 57,614.08 | 9.2 | 4 | 3 | −0.31 | Plasma membrane | |
| 24 | ZmAGC1.8 | LOC100281756 | 854 | 93,506.86 | 9.4 | 2 | 10 | −0.44 | Extracellular | |
| 25 | ZmAGC1.9 | LOC100381917 | 799 | 87,357.7 | 9.24 | 1 | 7 | −0.47 | Extracellular | |
| 26 | ZmAGC1.10 | LOC100273930 | 498 | 55,233.93 | 9.08 | 2 | 3 | −0.42 | Plasma membrane | |
| 27 | ZmAGC1.11 | LOC103633673 | 529 | 57,344.98 | 9.4 | 1 | 1 | −0.29 | Extracellular | |
| 28 | ZmAGC1.12 | LOC100192540 | 516 | 54,862.99 | 7.21 | 2 | 9 | −0.26 | Extracellular | |
| 29 | ZmPK3 | LOC103847826 | 530 | 57,414.92 | 9.38 | un | un | −0.33 | Extracellular | |
| 30 | ZmPK5 | LOC100274135 | 577 | 63,192.36 | 7.88 | 4 | 2 | −0.51 | Extracellular | |
| 31 | ZmPK7 | LOC100281711 | 583 | 63,635.37 | 8.36 | 3 | 4 | −0.4 | Plasma membrane | |
| 32 | ZmPID | LOC103643693 | 471 | 50,667.46 | 8.98 | 1 | 1 | −0.28 | Plasma membrane | |
| 33 | ZmKIPK | LOC103646509 | 799 | 87,303.65 | 9.27 | 2 | 2 | −0.47 | Extracellular | |
| 34 | ZmAGC2.1 | LOC107403167 | 455 | 48,371.7 | 6.82 | 2 | 2 | −0.27 | Extracellular | |
| 35 | ZmAGC2.2 | LOC100382773 | 467 | 49,785.03 | 6.24 | 2 | 10 | −0.32 | Plasma membrane | |
| 36 | ZmAGC2.3 | LOC100482936 | 438 | 47,503.72 | 6.29 | 4 | 8 | −0.25 | Extracellular | |
| 37 | ZmAGC2.4 | LOC100191446 | 438 | 47,493.65 | 6.33 | 1 | 2 | −0.28 | Plasma membrane | |
| 38 | ZmPHOT1 | LOC541663 | 911 | 101,471.3 | 8.66 | 22 | 3 | −0.47 | Plasma membrane | |
| 39 | ZmPHOT2 | LOC100281086 | 899 | 100,810.69 | 7.3 | 24 | 1 | −0.51 | Plasma membrane |
un, unknown.
Gene structures and motif analysis of AGC gene family in Z. mays
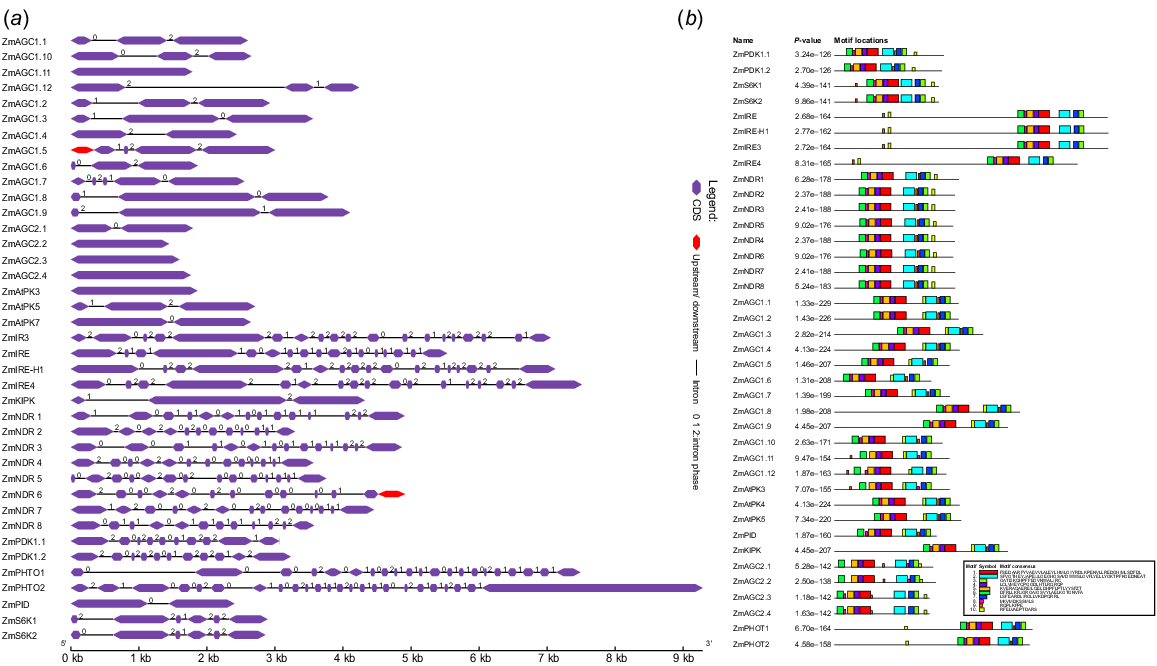
The intron and exon structures of ZmAGC members were analysed using the Gene Structure Display Server 2.0 to acquire a better knowledge of their structural properties. The study found that the number of introns in genes can vary (0–24). The ZmPHOT2 gene has the most introns, whereas ZmAGC2.2, ZmAGC2.3 and ZmAGC2.4 have no introns. Thus, the AGC1, AGC2 and related AGC and S6K gene families contain fewer intron and exon regions. Fig. 2b clearly shows that the site distribution of un-gapped motifs was chosen to be around 10. Motifs 1 and 2 were the most conserved across the entire ZmAGC kinase family. The motif logos was useful in predicting the functional motifs involved in AGC genes found in Z. mays. The site distribution of un-gapped motifs was chosen to be around 10. The motifs play an important role in performing their distinct function, and different colours are used in the figure to differentiate all of the conserved motifs as well as motif consensus.
Gene structure and motif analysis. (a) An online server Gene Structure Display Server 2.0 is used in which introns, exons, CDS, upstream and downstream sequences were emphasised. Coding sequence (CDS) regions are presented by purple bars, introns are represented by grey lines and upstream and downstream sequences are highlighted by red bars. The intron phase is shown by numbering 0, 1, 2. (b) Graphical representation of 10 conserved motifs found in all the protein sequences and their homologues. Each motif is represented by a distinct colour to differentiate it from the others. The presence and arrangement of these motifs are likely important for the function or structure of the proteins.
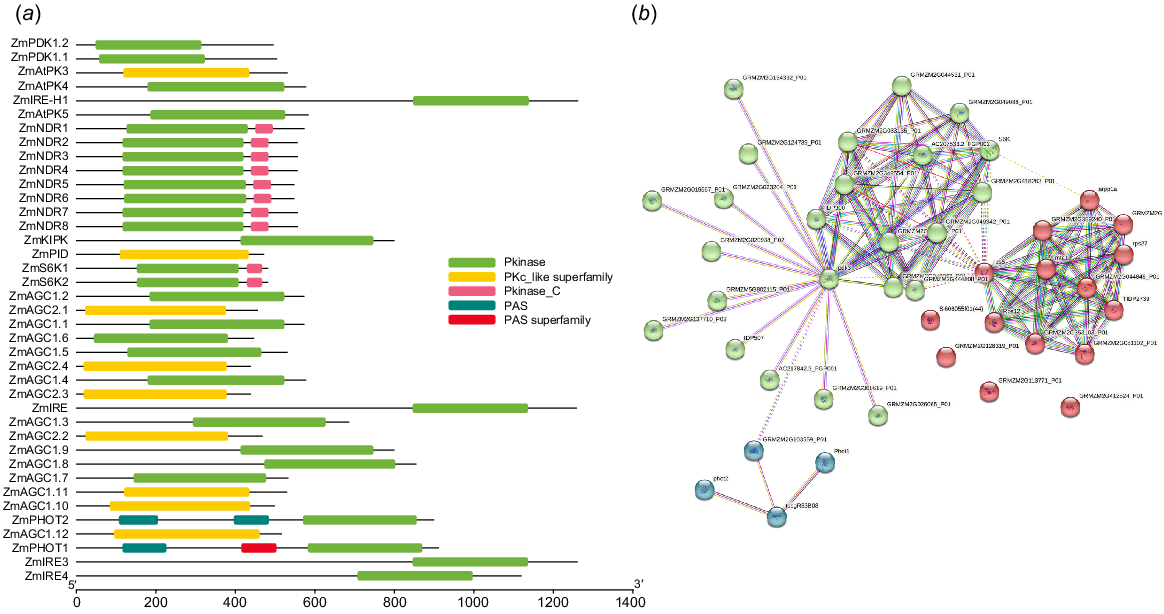
ZmAGC members domain analysis and protein-protein interaction analysis
Domain analysis of Z. mays revealed that members of the AGC gene family possessed conserved domains such as Pkinase, Pkinase_C and Per-Arnt-Sim (PAS). Pkinase or Pkinase_C was found in almost all proteins, whereas the PAS domain was found in only two proteins; (1) ZmHOT1; and (2) ZmHOT2. Protein–protein interactions revealed that members of the AGC gene family interacted with each other as well as with other proteins in the cell. ZmHOT1 and ZmHOT2 showed interaction with a discrete set of cellular proteins, while they showed weak interaction with the set of proteins that are members of the AGC gene family and other proteins with which they interact (Fig. 3).
Domain analysis and protein-protein interaction: (a) Protein domains of ZmAGCs were retrieved from NCBI CDD decamped. Pkinase is the most conserved domain. (b) Potein–protein interaction network was developed using the STRING database illustrating the most and least network of interacting proteins.
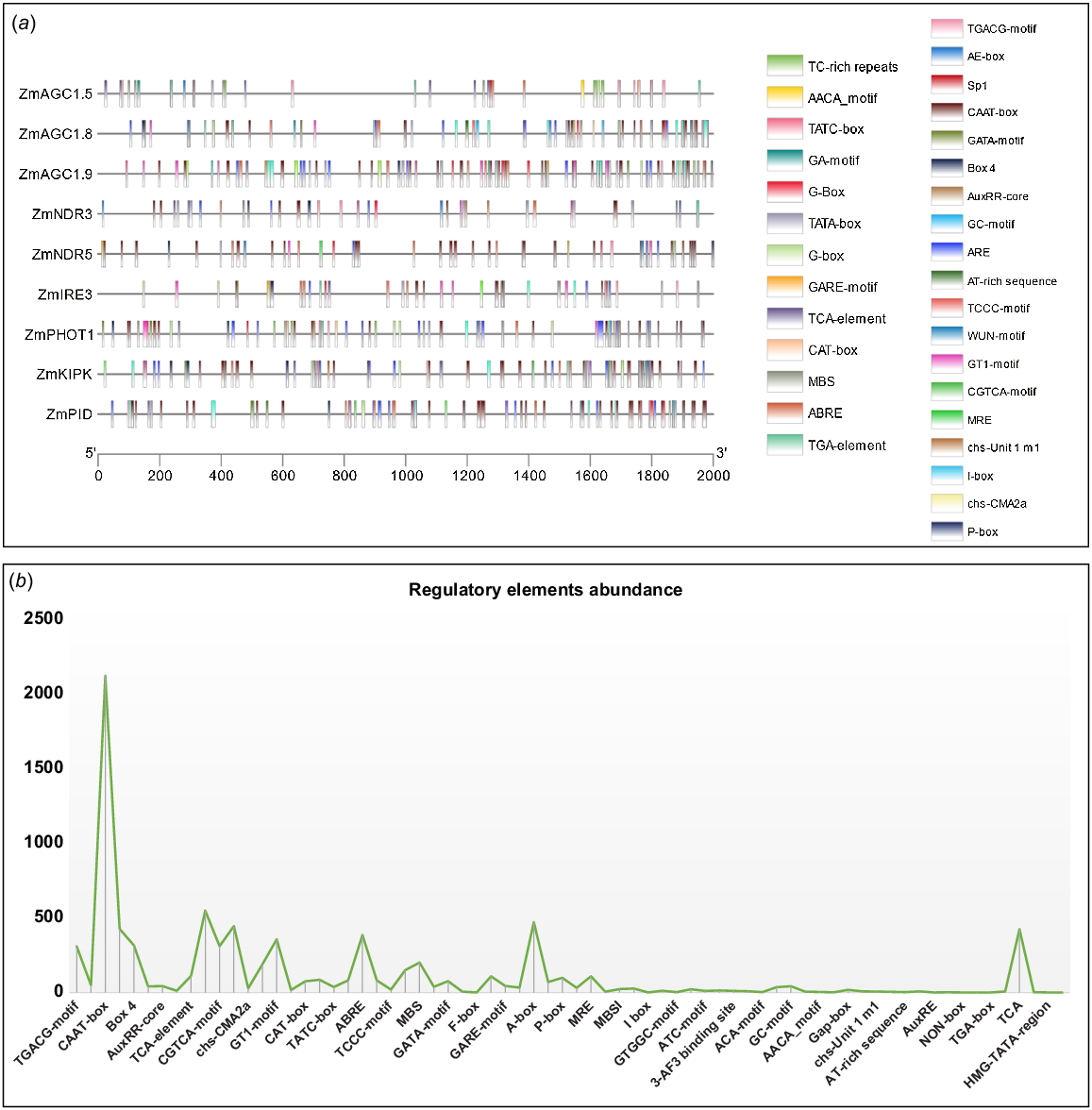
Promoter region and regulatory elements analysis of ZmAGC genes
The 2000-bp promoter sequences of 39 ZmAGC genes were examined for cis-regulatory elements. The analysis revealed 63 different types of regulatory elements, with the G-box being the most common. TGACG motif, AuxRR core, box-4, chs-CMA2a, 3-Af1 binding site, TC-rich repeats, AE-box, TATC-box, TCA, BOX-III and GAP-box were also common. Some elements were present in smaller numbers, such as B. HD Zip 1, GATT motif, GC motif, AT-rich element, WUN motif, LAMP elements, O2 site and I-Box. These results suggest that cis-regulatory elements play an important role in gene expression and shed light on the regulation of gene expression in Z. mays. Heat stress is mediated by TGACG motif, Sp1, CAAT box, G box, box 4, GCN4_Motif, AuxRR core, AT-rich element, TCA element, ARE, CGTCA motif and chs-CMA2a (Fig. 4).
Gene expression analysis of ZmAGC members under heat stress
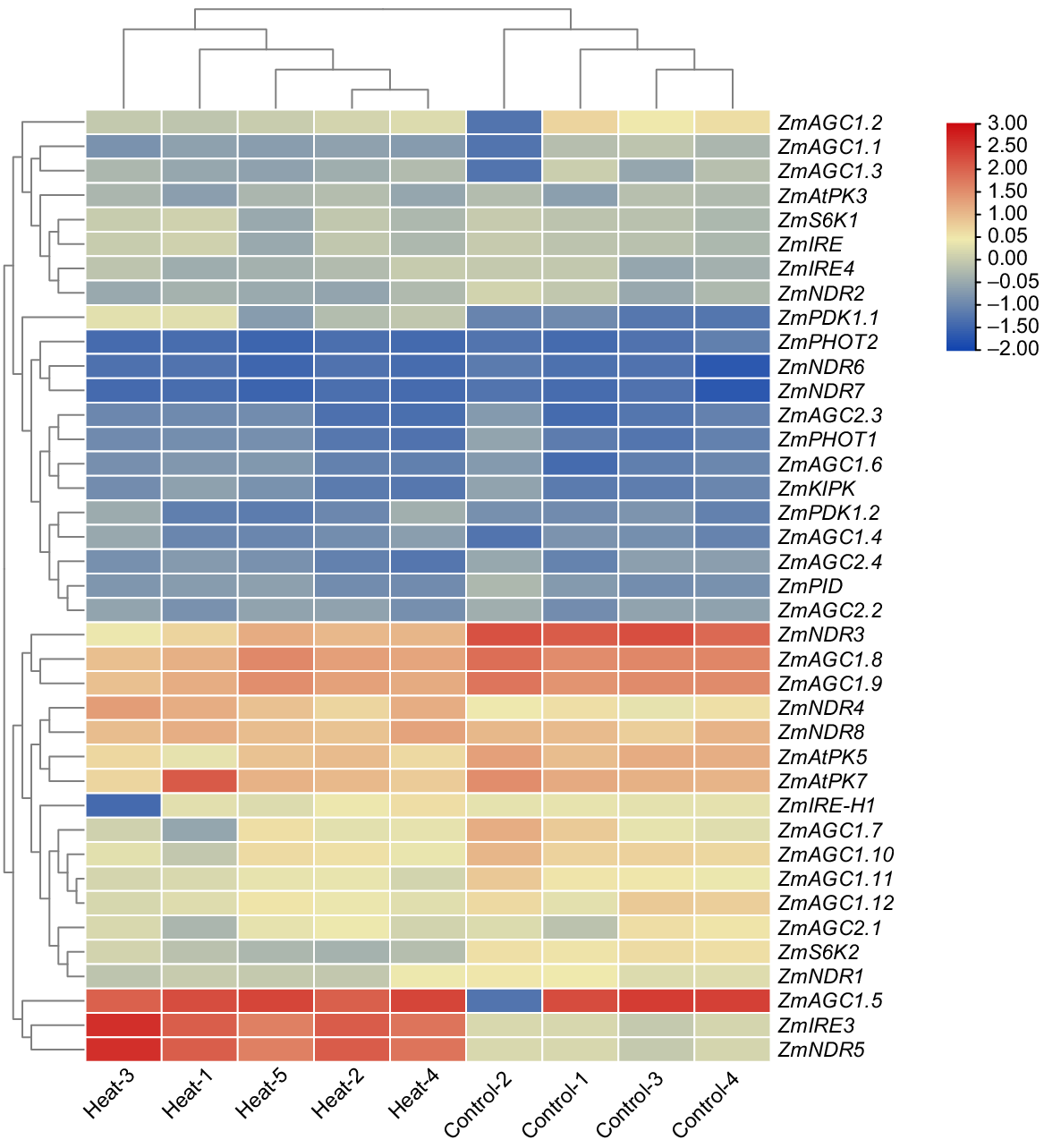
In this study, Next-Generation Sequencing (NGS) data analysis was employed to investigate the genetic response of Z. mays kernels under heat stress conditions. The research focused on identifying heat-related genes and unravelling the underlying biological pathways that have a significant impact on heat stress tolerance also potentially providing valuable insights for crop improvement strategies in the face of climate change. The expression profile of ZmAGC genes in Z. mays under heat stress (Fig. 5). The results showed that ZmNDR5 and ZmIRE3 were significantly upregulated (with at least a 2-fold change) in response to heat stress. ZmNDR4, on the other hand, showed a slight upregulation, but the change was not statistically significant. Furthermore, genes such as ZmAGC2.2, ZmPID, ZmKIPK, ZmIRE, ZmS6K1 and other members of this gene family did not show significant changes in expression, suggesting that they were not affected by heat stress. In contrast, ZmNDR3, ZmAGC1.8, ZmAGC1.9 and ZmAGC1.5 showed significant downregulation in response to heat stress with a fold change of 0.5 or less. Similarly, ZmNDR8, ZmAtPK5 and ZmAtPK7 showed slight downregulation, albeit not as pronounced. Taken together, these findings highlight the diverse response of ZmAGC genes to heat stress in Z. mays, with some genes significantly upregulated, others unchanged, and still others significantly downregulated.
Gene expression profiles of ZmAGCs are presented under the heat stress and heat map was generated using Tbtools and in addition to this, scaling is done between −2.50 and 3.00. Red, upregulated genes; blue, downregulated genes.
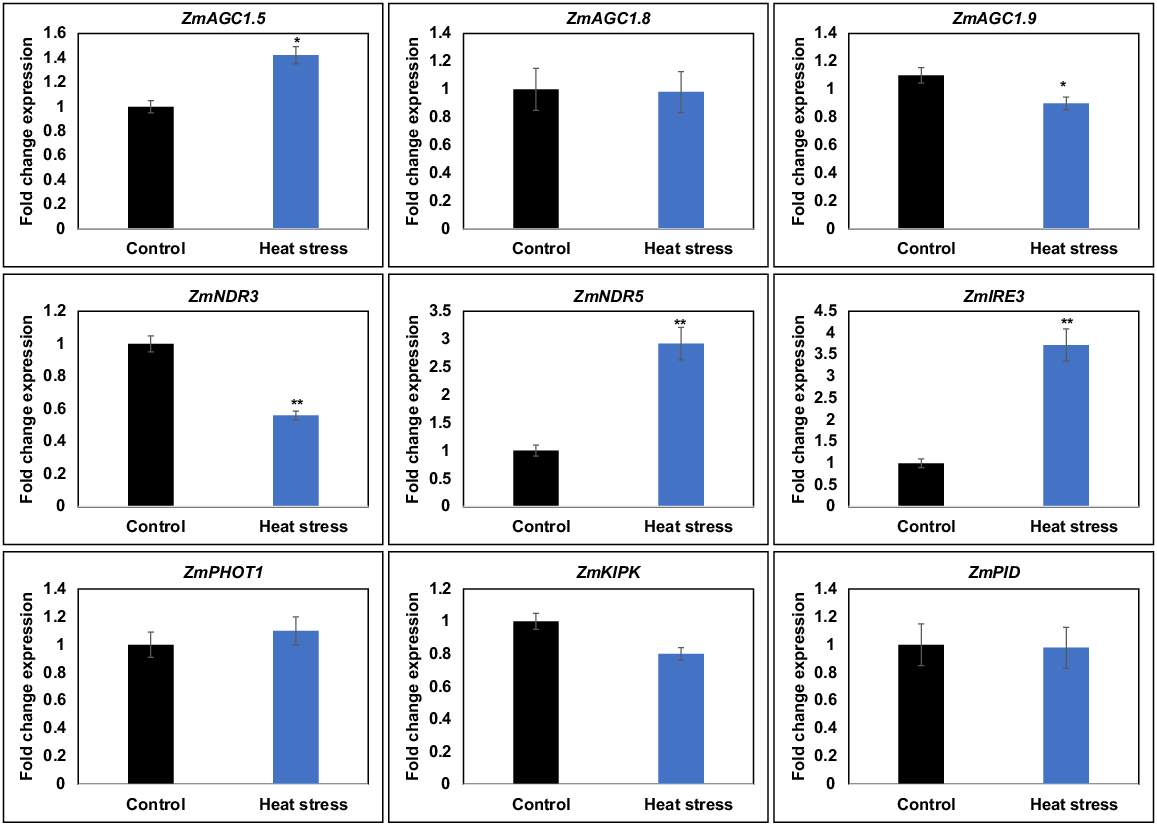
The gene transcript quantity of nine AGC genes in Z. mays kernels was determined using qPCR. Based on the NGS analysis the qPCR-based quantification of nine genes which include the upregulatory genes, downregulatory genes and genes that were identified as the PIN auxin efflux carriers were included like ZmAGC1.5, ZmIRE3, ZmNDR5, ZmNDR3, ZmAGC1.9, ZmPHOT1, ZmKIPK, ZmAGC1.8 and ZmPID was performed in response to heat. ZmAGC1.5 was found to be significantly expressed with heat stress, and genes such as ZmIRE3 and ZmNDR5 were strongly upregulated, while ZmNDR3 was significantly downregulated (Fig. 6).
Real-time qPCR analysis was conducted to investigate the expression of ZmAGCs in Z. mays leaves under heat stress conditions. The coloured bar graphs represent the mean expression levels ± s.e. based on three independent replicates. *, statistical significance (t-test) at the 95% confidence level; **, significance at the 99% confidence level.
Targeting of PIN auxin efflux carriers by AGC kinases in Z. mays
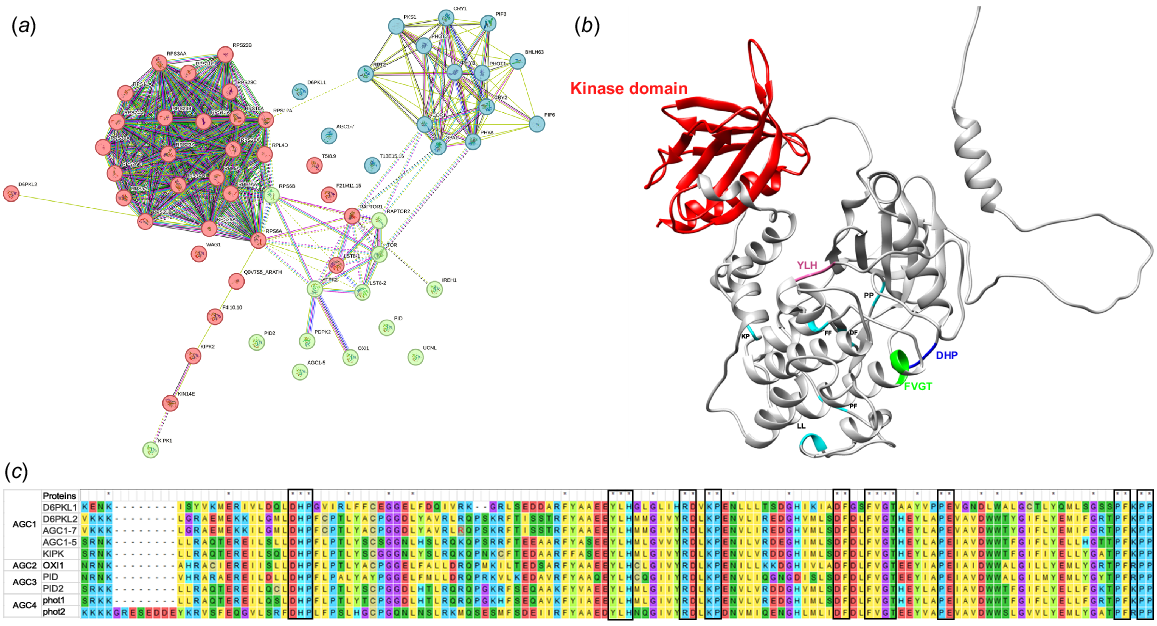
In the realm of vascular plants, the AGC1 to AGC4 kinases represent distinct subfamilies within the AGC kinase group. Interestingly, the evolutionary origins of all AGC kinases found in terrestrial plants can be traced back to PHOTOTROPIN2-like proteins originally present in algae. Fig. 7a illustrates the intricate network of protein–protein interactions within the PIN auxin factor proteins, further categorised into three distinct clusters that collectively house the Z. mays PIN auxin proteins. Within the vibrant Red Cluster, we observe the presence of AGC1 proteins, including D6PKL1, D6PKL2, AGC1-7 and AGC1-5. Shifting our attention to the Green Cluster, which belongs to AGC3, we find the protein members PID and PID2. Lastly, nestled in the Blue Cluster of AGC4, we encounter the Phot1 and Phot2 protein members. In Fig. 7b, we are presented with a detailed view of AGC1 kinase D6PKL1, allowing us to explore the intricate structure of this protein. Notably, the structure has been thoughtfully constructed using the Itasser server, and the consensus motif bases have been thoughtfully highlighted to enhance our understanding. The protein structure is colour-coded for clarity, with the red segments representing the kinase domain, while the green, blue and cyan regions denote the positions of the consensus motif bases. This comprehensive visualisation provides valuable insights into the key elements of this important protein. Fig. 7c, Notably, a consensus in the form of specific motifs has been identified within the sequences of all AGC family members, with common two-base amino acid pairs shared among some. These motifs, including DHP, YLH, and FVGT, along with pairs such as FF, PP, PF, DF and KP, provide intriguing insights into the functional diversity and evolutionary history of AGC kinases in Z. mays and beyond. In the case of Z. mays, two phototropin (phot) blue light receptor kinases are members of the AGC4 subfamily, playing pivotal roles in various light-responsive processes. Notably, they are deeply involved in regulating phototropic stem bending. For instance, Z. mays phot1 serves as a substantial adverse modulator of ATP-binding module B auxin transporters reduces auxin transport yet allows for transverse axis mobility during phototropic growth. Conversely, within Z. mays, the AGC1 kinase family, comprising 13 members, has diversified to acquire a range of regulatory functions in terrestrial plants. Remarkably, despite their evolutionary lineage from PHOTOTROPIN2-like proteins, these AGC1 kinases have retained their positions at the plasma membrane. Here, they play critical roles in auxin transportation apical development, enlargement of cells, and other activities. AGC3 kinases, on the other hand, have evolved to fulfil roles in controlling the polarity of PIN proteins, representing a critical adaptation to meet the developmental and differentiation needs of multicellular organisms. Meanwhile, AGC2 kinases have primarily been associated with nuclear functions and do not appear to be directly involved in phosphoregulatory processes.
(a) Protein-protein interaction network has been categorised into three distinct clusters: the red and blue clusters encompass AGC1 members; while the green cluster emcompasses the AGC2 and AGC3 members. AGC4 members are represented within the blue cluster. (b) We present a model of the AGC1 kinase D6PKL1, showcasing the protein’s structure. The consensus motif bases have been thoughtfully highlighted in the structure, which was constructed using the Itasser server. (c) In the protein sequence alignment of the insertion loop of selected Z. mays AGC kinases, various sequence features have been marked. Notably, the sequence variation within the DHP motif may hold functional significance, particularly for D6PKL1, KIPK and D6PKL2, which are proposed as AGC1 targets.
Discussion
Maize has been extensively investigated as a model plant, and with the in-depth sequencing of its genome, there has been a refinement in the details of genome annotation. This comprehensive sequencing has enabled the identification of crucial genes associated with growth and development. In recent studies, several gene families, including NHL, NF-Y and CAMTA, have been pinpointed in maize (Cao et al. 2023; Liu et al. 2023; Shahbaz et al. 2023). The AGC protein kinase family, a subset of the broader protein kinase superfamily, plays a pivotal role in regulating plant growth and development (Jiang et al. 2022; Wu et al. 2022). The initial discovery of the AGC gene family took place in A. thaliana, leading to numerous functional investigations. These studies confirmed the involvement of AGC protein kinases in diverse processes such as lipid signalling pathways, auxin regulation and cell proliferation (Bögre et al. 2003). While functional studies of AGC genes in Triticum aestivum, Solanum lycopersicum, Pyropia yezoensis and Medicago truncatula have been limited, they have shown associations with apoptosis, defence response and cell division (Pislariu and Dickstein 2007; Ek-Ramos et al. 2010; Zhu et al. 2015, 2022; Liu et al. 2017), despite the absence of systematic identification of AGC genes in these plants. As of now, the distribution of AGC genes in maize remains unknown, This study systematically identified AGC genes in maize, conducting analyses on their characterisation, sequence structure, motifs, phylogenetics and cis-acting elements. Furthermore, the expression profiles of AGC genes were scrutinised through a combination of RNA-seq and real-time qPCR to provide a comprehensive understanding of their functions.
Genome-wide analysis of AGC gene family in Z. mays resulted into several intriguing features. The AGC gene family in Z. mays is incredibly diverse, with notable variations in gene number, structure, and expression patterns within subfamilies (Table 1).
Genome-wide analysis of AGC gene family in Z. mays resulted in several intriguing features. The AGC gene family in Z. mays is incredibly diverse, with notable variations in gene number, structure, and expression patterns within subfamilies (Table 1). The number of subfamilies and members in these subfamilies were consistent with A. thaliana. Such conservation of family members has been reported for other gene families like MBD gene family for Z. mays and A. thaliana (Qian et al. 2020). This consistency implies a degree of evolutionary conservation in the AGC kinase gene family across these plant species, highlighting the importance of these kinases in fundamental cellular processes that have been maintained throughout evolution (Qian et al. 2020).
The physiochemical properties of AGC kinase subfamilies in maize reveal diverse characteristics that likely contribute to their functional roles. Analysing the protein features, it is evident that the subfamilies exhibit variations in amino acid length, molecular weight, theoretical isoelectric point (pI) and GRAVY (grand average of hydropathy) values. For instance, ZmIRE and ZmNDR subfamilies possess considerably larger proteins with higher molecular weights, potentially indicating complex structural configurations associated with their roles. However, ZmAGC1 and ZmAGC2 subfamilies show a range of molecular weights, suggesting functional diversity within these groups. The theoretical isoelectric points vary across the subfamilies, reflecting differences in the net charge of the proteins under physiological conditions. The GRAVY values indicate the hydrophobic or hydrophilic nature of the proteins, with subfamilies like ZmNDR exhibiting relatively hydrophobic characteristics. These physiochemical variations likely contribute to the functional diversity observed within the AGC kinase subfamilies in maize, suggesting specific roles and interactions in various cellular processes. The physiochemical properties observed in AGC kinase subfamilies in maize share parallels with findings in other gene families. For instance, studies on the NAC transcription factor family in maize have highlighted the significance of diverse protein lengths and molecular weights, impacting their functions in developmental processes (Wang et al. 2020). Similarly, investigations into the WRKY transcription factor family have demonstrated variations in isoelectric points and GRAVY values, emphasising their roles in stress responses (Hu et al. 2021). These examples underscore the broader patterns of physiochemical diversity within maize gene families, supporting the notion that such variations are likely crucial for their specific functions in cellular processes.
The prediction of subcellular protein localisation is a crucial aspect of understanding the functions of proteins within a cell. Kinases involved in cellular signalling (ZmPDK1.1, ZmPDK1.2, ZmS6K1, ZmS6K2 in cytoplasm/plasma membrane) are often found in the cytoplasm where they can relay signals. Similarly, Proteins localised to the nucleus (ZmIRE, ZmIRE-H1, ZmIRE3, ZmIRE4, ZmNDR2, ZmNDR3, ZmNDR5, ZmNDR6, ZmNDR7, ZmNDR8) are frequently associated with transcriptional regulation, DNA replication and other nuclear processes. NDR kinases, for instance, are known to play roles in cell cycle control and may exert their functions in the nucleus (Yoon et al. 2021). Extracellular proteins are typically involved in cell communication, signalling, or response to environmental stimuli. Kinases like ZmPK3, ZmPK5 and ZmPK7, as well as receptor-like proteins (e.g. ZmPID), may interact with extracellular ligands to transmit signals (Almagro et al. 2009). AGC kinases and phototropins (ZmAGC1.3, ZmAGC1.4, ZmAGC1.6, ZmAGC1.7, ZmAGC1.10, ZmAGC2.2, ZmAGC2.4, ZmPHOT1, ZmPHOT2) are known to localise to the plasma membrane, where they can interact with other membrane-associated proteins (Hirano et al. 2022).
Motifs within Class III peroxidases offers valuable insights into their functional significance. One common motif is the heme-binding domain, crucial for peroxidase activity as it facilitates the binding of haem, a cofactor central to the enzyme’s catalytic function in oxidative reactions. Another important motif is the conserved amino acid residues responsible for substrate binding and specificity, influencing the types of molecules the peroxidase can interact with and modify (Mathé et al. 2010) SnRK1 (sucrose non-fermenting 1-related protein kinase 1) and SnRK2 (sucrose non-fermenting 1-related protein kinase 2), which are implicated in energy and stress signalling pathways, respectively, are well-characterised genes found in ZmAGC Z. mays members (Kulik et al. 2011). Numerous biological processes, such as stress reactions, hormone signalling, metabolism, and development, are influenced by the Z. mays AGC gene family (Ehlting et al. 2008).
The identification of cis-regulatory elements within the 2000-bp promoter sequences of 39 ZmAGC genes provides critical insights into the regulation of gene expression, particularly in the context of heat stress (Ijaz et al. 2020). The prevalence of the G-box, TGACG motif, AuxRR core, box-4, chs-CMA2a and other elements suggests their significant role in orchestrating the transcriptional response to environmental stimuli, especially heat stress. The TGACG motif, known as the cis-acting element involved in the MeJA (methyl jasmonate) responsiveness, indicates a potential link between ZmAGC genes and stress signalling pathways. Elements like Sp1, CAAT box, and G box are associated with general transcriptional regulation, while GCN4_Motif is linked to amino acid biosynthesis and stress response. The presence of AT-rich elements, TCA elements and ARE points towards their involvement in abiotic stress responses, including heat stress. The observed differential expression of ZmAGC genes under heat stress in Z. mays reveals potential functional implications and provides valuable insights into the molecular mechanisms governing the plant’s response to elevated temperatures (Abeed et al. 2022; Eljebbawi et al. 2022; Ahmed et al. 2023). Firstly, the significant upregulation of ZmNDR5 and ZmIRE3 suggests a positive role for these genes in the heat stress response. These genes may encode proteins that contribute to stress tolerance, possibly through involvement in signalling pathways or molecular processes that help the plant cope with thermal stress (Mathé et al. 2010; Abeed et al. 2021, 2022). On the contrary, the downregulation of genes like ZmNDR3, ZmAGC1.8, ZmAGC1.9 and ZmAGC1.5 indicates a negative impact on their expression levels, potentially affecting their normal functions. ZmAGC1.5, in particular, showed significant upregulation, emphasising its potential role as a key player in the plant’s heat stress response. Real-time qPCR results corroborate the NGS findings, further validating the differential expression patterns of these genes under heat stress conditions. The inclusion of PIN auxin efflux carriers in the analysis suggests a potential link between AGC genes and auxin signalling, indicating a complex regulatory network responding to heat stress (Chen et al. 2022). Overall, these findings contribute to our understanding of the molecular basis of heat stress tolerance in Z. mays and may guide future efforts in crop improvement for enhanced resilience to climate change-induced challenges.
Declaration of funding
Princess Nourah bint Abdulrahman University Researchers Supporting Project number (PNURSP2024R402), Princess Nourah bint Abdulrahman University, Riyadh, Saudi Arabia.
References
Abeed AHA, Ali M, Ali EF, Majrashi A, Eissa MA (2021) Induction of Catharanthus roseus secondary metabolites when Calotropis procera was used as bio-stimulant. Plants 10, 1623.
| Crossref | Google Scholar | PubMed |
Abeed AHA, Ali M, Eissa MA, Tammam SA (2022) Impact of sewage water irrigation on Datura innoxia grown in sandy loam soil. BMC Plant Biology 22, 559.
| Crossref | Google Scholar |
Ahmed T, Masood HA, Noman M, Al-Huqail AA, Alghanem SM, Khan MM, Muhammad S, Manzoor N, Rizwan M, Qi X, Abeed AHA, Li B (2023) Biogenic silicon nanoparticles mitigate cadmium (Cd) toxicity in rapeseed (Brassica napus L.) by modulating the cellular oxidative stress metabolism and reducing Cd translocation. Journal of Hazardous Materials 459, 132070.
| Crossref | Google Scholar | PubMed |
Almagro L, Gómez Ros LV, Belchi-Navarro S, Bru R, Ros Barceló A, Pedreño MA (2009) Class III peroxidases in plant defence reactions. Journal of Experimental Botany 60, 377-390.
| Crossref | Google Scholar | PubMed |
Anthony RG, Henriques R, Helfer A, Mészáros T, Rios G, Testerink C, Munnik T, Deák M, Koncz C, Bögre L (2004) A protein kinase target of a PDK1 signalling pathway is involved in root hair growth in Arabidopsis. The EMBO Journal 23, 572-581.
| Crossref | Google Scholar | PubMed |
Bailey TL, Johnson J, Grant CE, Noble WS (2015) The MEME suite. Nucleic Acids Research 43, W39-W49.
| Crossref | Google Scholar | PubMed |
Benson DA, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (2010) GenBank. Nucleic Acids Research 38, D46-D51.
| Crossref | Google Scholar |
Bigini V, Camerlengo F, Botticella E, Sestili F, Savatin DV (2021) Biotechnological resources to increase disease-resistance by improving plant immunity: a sustainable approach to save cereal crop production. Plants 10, 1146.
| Crossref | Google Scholar | PubMed |
Biondi RM (2004) Phosphoinositide-dependent protein kinase 1, a sensor of protein conformation. Trends in Biochemical Sciences 29, 136-142.
| Crossref | Google Scholar | PubMed |
Bögre L, Ökrész L, Henriques R, Anthony RG (2003) Growth signalling pathways in Arabidopsis and the AGC protein kinases. Trends in Plant Science 8, 424-431.
| Crossref | Google Scholar | PubMed |
Cao L, Ma C, Ye F, Pang Y, Wang G, Fahim AM, Lu X (2023) Genome-wide identification of NF-Y gene family in maize (Zea mays L.) and the positive role of ZmNF-YC12 in drought resistance and recovery ability. Frontiers in Plant Science 14, 1159955.
| Crossref | Google Scholar | PubMed |
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Molecular Plant 13, 1194-1202.
| Crossref | Google Scholar | PubMed |
Chen L, Cai M, Chen M, Ke W, Pan Y, Huang J, Zhang J, Peng C (2022) Genome-wide characterization of PIN auxin efflux carrier gene family in Mikania micrantha. International Journal of Molecular Sciences 23, 10183.
| Crossref | Google Scholar | PubMed |
Crafts-Brandner SJ, Salvucci ME (2002) Sensitivity of photosynthesis in a C4 plant, maize, to heat stress. Plant Physiology 129, 1773-1780.
| Crossref | Google Scholar | PubMed |
Dhonukshe P, Huang F, Galvan-Ampudia CS, Mähönen AP, Kleine-Vehn J, Xu J, Quint A, Prasad K, Friml J, Scheres B, Offringa R (2010) Plasma membrane-bound AGC3 kinases phosphorylate PIN auxin carriers at TPRXS(N/S) motifs to direct apical PIN recycling. Development 137, 3245-3255.
| Crossref | Google Scholar | PubMed |
Dubé N, Tremblay ML (2005) Involvement of the small protein tyrosine phosphatases TC-PTP and PTP1B in signal transduction and diseases: from diabetes, obesity to cell cycle, and cancer. Biochimica et Biophysica Acta (BBA) - Proteins and Proteomics 1754, 108-117.
| Crossref | Google Scholar | PubMed |
Ehlting J, Chowrira SG, Mattheus N, Aeschliman DS, Arimura G-I, Bohlmann J (2008) Comparative transcriptome analysis of Arabidopsis thaliana infested by diamond back moth (Plutella xylostella) larvae reveals signatures of stress response, secondary metabolism, and signalling. BMC Genomics 9, 154.
| Crossref | Google Scholar |
Ek-Ramos MJ, Avila J, Cheng C, Martin GB, Devarenne TP (2010) The T-loop extension of the tomato protein kinase AvrPto-dependent Pto-interacting protein 3 (Adi3) directs nuclear localization for suppression of plant cell death. Journal of Biological Chemistry 285, 17584-17594.
| Crossref | Google Scholar | PubMed |
Eljebbawi A, Savelli B, Libourel C, Estevez JM, Dunand C (2022) Class III peroxidases in response to multiple abiotic stresses in Arabidopsis thaliana pyrenean populations. International Journal of Molecular Sciences 23, 3960.
| Crossref | Google Scholar | PubMed |
Galván-Ampudia CS, Offringa R (2007) Plant evolution: AGC kinases tell the auxin tale. Trends in Plant Science 12(12), 41-47.
| Crossref | Google Scholar |
Garcia AV, Al-Yousif M, Hirt H (2012) Role of AGC kinases in plant growth and stress responses. Cellular and Molecular Life Sciences 69, 3259-3267.
| Crossref | Google Scholar | PubMed |
Garg VK, Avashthi H, Tiwari A, Jain PA, Ramkete PW, Kayastha AM, Singh VK (2016) MFPPI–multi FASTA ProtParam interface. Bioinformation 12, 74-77.
| Crossref | Google Scholar | PubMed |
Garnacho-Montero J, Amaya-Villar R (2010) Multiresistant Acinetobacter baumannii infections: epidemiology and management. Current Opinion in Infectious Diseases 23, 332-339.
| Crossref | Google Scholar | PubMed |
Geer LY, Marchler-Bauer A, Geer RC, Han L, He J, He S, Liu C, Shi W, Bryant SH (2010) The NCBI biosystems database. Nucleic Acids Research 38, D492-D496.
| Crossref | Google Scholar | PubMed |
Hajibarat Z, Saidi A (2022) Genome wide identification of AGC kinase genes and their expression in response to heat and cold stresses in barley. Acta Agriculturae Slovenica 118, 1-11.
| Crossref | Google Scholar |
Harris D, Breese WA, Rao JVDKK (2005) The improvement of crop yield in marginal environments using ‘on-farm’ seed priming: nodulation, nitrogen fixation, and disease resistance. Australian Journal of Agricultural Research 56, 1211-1218.
| Crossref | Google Scholar |
Hirano S, Sasaki K, Osaki Y, Tahara K, Takahashi H, Takemiya A, Kodama Y (2022) The localization of phototropin to the plasma membrane defines a cold-sensing compartment in Marchantia polymorpha. PNAS Nexus 1, pgac030.
| Crossref | Google Scholar |
Howe LR, Leevers SJ, Gómez N, Nakielny S, Cohen P, Marshall CJ (1992) Activation of the MAP kinase pathway by the protein kinase raf. Cell 71, 335-342.
| Crossref | Google Scholar | PubMed |
Hu W, Ren Q, Chen Y, Xu G, Qian Y (2021) Genome-wide identification and analysis of WRKY gene family in maize provide insights into regulatory network in response to abiotic stresses. BMC Plant Biology 21, 427.
| Crossref | Google Scholar |
Ijaz U, Pervaiz T, Ahmed T, Seemab R, Shahid M, Noman M, Nadeem M (2020) Plant Cis-regulatory elements: methods of identification and applications. Asian Journal of Agriculture and Biology 8, 207-222.
| Crossref | Google Scholar |
Jiang Y, Liu X, Zhou M, Yang J, Ke S, Li Y (2022) Genome-wide identification of the AGC protein kinase gene family related to photosynthesis in rice (Oryza sativa). International Journal of Molecular Sciences 23, 12557.
| Crossref | Google Scholar | PubMed |
Kibbe WA (2007) OligoCalc: an online oligonucleotide properties calculator. Nucleic Acids Research 35, W43-W46.
| Crossref | Google Scholar | PubMed |
Kulik A, Wawer I, Krzywińska E, Bucholc M, Dobrowolska G (2011) SnRK2 protein kinases—key regulators of plant response to abiotic stresses. Omics: a Journal of Integrative Biology 15, 859-872.
| Crossref | Google Scholar | PubMed |
Leroux AE, Schulze JO, Biondi RM (2018) AGC kinases, mechanisms of regulation and innovative drug development. Seminars in Cancer Biology 48, 1-17.
| Crossref | Google Scholar | PubMed |
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Research 30, 325-327.
| Crossref | Google Scholar | PubMed |
Letunic I, Bork P (2019) Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Research 47, W256-W259.
| Crossref | Google Scholar | PubMed |
Liu J-Z, Duan J, Ni M, Liu Z, Qiu W-L, Whitham SA, Qian W-J (2017) S-Nitrosylation inhibits the kinase activity of tomato phosphoinositide-dependent kinase 1 (PDK1). Journal of Biological Chemistry 292, 19743-19751.
| Crossref | Google Scholar | PubMed |
Liu W, Du H, Huang M (2023) Genome-wide analysis of the CAMTA gene family in maize (Zea mays L.). Maize Genomics and Genetics 14, 1-8.
| Crossref | Google Scholar |
Marchler-Bauer A, Derbyshire MK, Gonzales NR, Lu S, Chitsaz F, Geer LY, Geer RC, He J, Gwadz M, Hurwitz DI, Lanczycki CJ, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Bryant SH (2015) CDD: NCBI’s conserved domain database. Nucleic Acids Research 43, D222-D226.
| Crossref | Google Scholar | PubMed |
Mathé C, Barre A, Jourda C, Dunand C (2010) Evolution and expression of class III peroxidases. Archives of Biochemistry and Biophysics 500, 58-65.
| Crossref | Google Scholar | PubMed |
Mering Cv, Huynen M, Jaeggi D, Schmidt S, Bork P, Snel B (2003) STRING: a database of predicted functional associations between proteins. Nucleic Acids Research 31, 258-261.
| Crossref | Google Scholar |
Mora A, Komander D, van Aalten DMF, Alessi DR (2004) PDK1, the master regulator of AGC kinase signal transduction. Seminars in Cell & Developmental Biology 15, 161-170.
| Crossref | Google Scholar | PubMed |
Nguyen L-T, Schmidt HA, Von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular Biology and Evolution 32, 268-274.
| Crossref | Google Scholar | PubMed |
Pearce LR, Komander D, Alessi DR (2010) The nuts and bolts of AGC protein kinases. Nature Reviews Molecular Cell Biology 11, 9-22.
| Crossref | Google Scholar | PubMed |
Pislariu CI, Dickstein R (2007) An IRE-like AGC kinase gene, MtIRE, has unique expression in the invasion zone of developing root nodules in Medicago truncatula. Plant Physiology 144, 682-694.
| Crossref | Google Scholar | PubMed |
Qian Y, Ren Q, Jiang L, Zhang J, Chen C, Chen L (2020) Genome-wide analysis of maize MBD gene family and expression profiling under abiotic stress treatment at the seedling stage. Plant Biotechnology Reports 14, 323-338.
| Crossref | Google Scholar |
Shahbaz M, Azeem F, Rafique MU, Siraj HMS, Rizwan M (2023) Heat-induced transcriptome and genome-wide analysis of NHL genes in maize (Zea mays L.) suggest a role of ZmNHLs under heat stress. Journal of Plant Growth Regulation 42, 6891-6902.
| Crossref | Google Scholar |
Tamaskovic R, Bichsel SJ, Hemmings BA (2003) NDR family of AGC kinases – essential regulators of the cell cycle and morphogenesis. FEBS Letters 546, 73-80.
| Crossref | Google Scholar | PubMed |
Wang G, Yuan Z, Zhang P, Liu Z, Wang T, Wei L (2020) Genome-wide analysis of NAC transcription factor family in maize under drought stress and rewatering. Physiology and Molecular Biology of Plants 26, 705-717.
| Crossref | Google Scholar | PubMed |
Wilhelm EP, Mullen RE, Keeling PL, Singletary GW (1999) Heat stress during grain filling in maize: effects on kernel growth and metabolism. Crop Science 39, 1733-1741.
| Crossref | Google Scholar |
Wu M-H, Yu Q, Tao T-Y, Sun L-X, Qian H, Zhu X-M, Li L, Liang S, Lu J-P, Lin F-C, Liu X-H (2022) Genome-wide analysis of AGC kinases reveals that MoFpk1 is required for development, lipid metabolism, and autophagy in hyperosmotic stress of the rice blast fungus Magnaporthe oryzae. mBio 13, e02279–22.
| Crossref | Google Scholar | PubMed |
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 13, 134.
| Crossref | Google Scholar |
Yoon HS, Fujino K, Liu S, Takano T, Tsugama D (2021) NDR/LATS-family protein kinase genes are indispensable for embryogenesis in Arabidopsis. FEBS Open Bio 11, 2600-2606.
| Crossref | Google Scholar | PubMed |
Zegzouti H, Li W, Lorenz TC, Xie M, Payne CT, Smith K, Glenny S, Payne GS, Christensen SK (2006) Structural and functional insights into the regulation of Arabidopsis AGC VIIIa kinases. Journal of Biological Chemistry 281, 35520-35530.
| Crossref | Google Scholar | PubMed |
Zhou P, Liang Y, Mei J, Liao H, Wang P, Hu K, Chen L, Zhang X, Ye D (2021) The Arabidopsis AGC kinases NDR2/4/5 interact with MOB1A/1B and play important roles in pollen development and germination. The Plant Journal 105, 1035-1052.
| Crossref | Google Scholar |
Zhu X, Yang K, Wei X, Zhang Q, Rong W, Du L, Ye X, Qi L, Zhang Z (2015) The wheat AGC kinase TaAGC1 is a positive contributor to host resistance to the necrotrophic pathogen Rhizoctonia cerealis. Journal of Experimental Botany 66, 6591-6603.
| Crossref | Google Scholar | PubMed |
Zhu Y, Wang X, Sun B, Tang X, Mao Y (2022) Cytological and transcriptional analysis reveal phosphatidylinositol signaling pathway plays key role in mitotic division of Pyropia yezoensis. Journal of Oceanology and Limnology 40, 1148-1159.
| Crossref | Google Scholar |