Functional characterisation of Dof gene family and expression analysis under abiotic stresses and melatonin-mediated tolerance in pitaya (Selenicereus undatus)
Osama Alam A # , Latif Ullah Khan B C # , Adeel Khan A , Saleh H. Salmen D , Mohammad Javed Ansari E , Fizza Mehwish A , Mushtaq Ahmad A , Qamar U. Zaman
A # , Latif Ullah Khan B C # , Adeel Khan A , Saleh H. Salmen D , Mohammad Javed Ansari E , Fizza Mehwish A , Mushtaq Ahmad A , Qamar U. Zaman  B C * and Hua-Feng Wang
B C * and Hua-Feng Wang  B C *
B C *
A
B
C
D
E
Handling Editor: Muhammad Nadeem
Abstract
DNA binding proteins with one finger (Dof) transcription factors are essential for seed development and defence against various biotic and abiotic stresses in plants. Genomic analysis of Dof has not been determined yet in pitaya (Selenicereus undatus). In this study, we have identified 26 Dof gene family members, renamed as HuDof-1 to HuDof-26, and clustered them into seven subfamilies based on conserved motifs, domains, and phylogenetic analysis. The gene pairs of Dof family members were duplicated by segmental duplications that faced purifying selection, as indicated by the Ka/Ks ratio values. Promoter regions of HuDof genes contain many cis-acting elements related to phytohormones including abscisic acid, jasmonic acid, gibberellin, temperature, and light. We exposed pitaya plants to different environmental stresses and examined melatonin’s influence on Dof gene expression levels. Signifcant expression of HuDof-2 and HuDof-6 were observed in different developmental stages of flower buds, flowers, pericarp, and pulp. Pitaya plants were subjected to abiotic stresses, and transcriptome analysis was carried out to identify the role of Dof gene family members. RNA-sequencing data and reverse transcription quantitative PCR-based expression analysis revealed three putative candidate genes (HuDof-1, HuDof-2, and HuDof-8), which might have diverse roles against the abiotic stresses. Our study provides a theoretical foundation for functional analysis through traditional and modern biotechnological tools for pitaya trait improvement.
Keywords: abiotic stress tolerance, climate resilience, Dof gene family, dragon fruit (Selenicereus undatus L.), melatonin-mediated tolerance, pitaya, RT-qPCR-based expression analysis, transcriptome analysis.
Introduction
Pitaya (Selenicereus undatus) is a tropical fruit known as dragon fruit. It belongs to the Cactaceae family, which has 127 genera and 1750 species (Zaman et al. 2023a). The pitaya plant is a species of the Hylocereus genus. In the recent past, the Hylocereus genus was transferred to the Selenicereus genus (Ruiz-Domínguez et al. 2019). It is a perennial diploid (2n = 22) that is native to Mexico and grown in all tropical and subtropical regions (Mercado-Silva 2018). The fruit is rich in vitamin C, phenolics, and betaine compounds and has strong antioxidant properties (Paśko et al. 2021).
Plant transcription factors (TFs) are proteins that participate in the regulation of genes by binding to DNA sequences in regulatory regions. They help to turn the gene on and off in response to various signals (Todeschini et al. 2014). The Dof protein is a TF that is abundant in nature and belongs to the zinc finger protein family. It was discovered in plants and has not yet been studied in other eukaryotes, such as yeast, protists, and mammals (Gupta et al. 2015). Dof proteins consist of 200–400 conserved amino acids; the N-terminus of the domain contains 50–52 amino acids (Yanagisawa and Sheen 1998). It contains a transcriptional regulatory region with a highly variable nuclear localisation signal at the C-terminus (Yanagisawa 2002). In the promoter region, the target gene contains an AAAG cis-element that is bounded by the Cys2/Cys2 zinc finger structure, which is present in the Dof domain and assumed to be a dual domain with important functions in binding to DNA and interaction of protein (Umemura et al. 2004). ZmDof1, the first Dof protein, was initially thought to be a DNA-binding protein when it was cloned from maize plants (Yanagisawa and Schmidt 1999). The number of Dof genes in species varies. For example, Arabidopsis thaliana is comprised of 36 gene family members, rice (Oryza sativa) contains 30 (Lijavetzky et al. 2003), and there are 26 in barley (Hordeum vulgare) (Moreno-Risueno et al. 2007), 46 in maize (Zea mays) (Chen and Cao 2015), 28 in sorghum (Sorghum bicolor) (Kushwaha et al. 2011), 36 in tomato (Solanum lycopersicum) (Cai et al. 2013), and 96 in wheat (Triticum aestivum) (Liu et al. 2020). A few researchers have recognised the distinct function of Dof proteins in numerous biological systems including metabolic control, seedling growth, and tissue differentiation (Noguero et al. 2013), complex vascular formation (Konishi and Yanagisawa 2007), blooming season (Negi et al. 2013), pollen development (Yang et al. 2011), germination (Gualberti et al. 2002), endosperm formation (Marzábal et al. 2008), metabolism of carbon and nitrogen (Santos et al. 2012), and responses to phytohormones (Venkatesh and Park 2015; Su et al. 2017).
In this analysis, we performed a genome-wide study of the Dof gene family in pitaya genome. We identified 26 Dof family members, which were mapped over 11 chromosomes. Based on RNA sequencing (RNA-seq) analysis, we selected a few genes differentially expressing under single-factor, double-factor, and multifactorial stresses. From our results, we have identified three candidate genes, that might have diverse role among Dof gene family members, playing diverse role against the abiotic stresses. Our identified putative candidate genes in pitaya can aid breeders in conventional and modern breeding technologies including CRISPR system (clustered regularly interspaced short palindromic repeats).
Materials and methods
Characterisation of the Dof gene family members in pitaya
To identify the Dof gene family members, we used the pitaya genome database (http://www.pitayagenomic.com, accessed on 13 March 2023) (Chen et al. 2022a). Dof family members of maize, rice, and Arabidopsis were retrieved from Phytozome (https://phytozome-next.jgi.doe.gov, accessed on 17 October 2023) (Goodstein et al. 2012) by blasting Dof sequence. To verify the domains of Dof proteins, we used the InterPro tool (https://www.ebi.ac.uk/interpro, accessed on 13 March 2023). Additionally, for the physiological and chemical characteristics of Dof proteins, we calculated their molecular weight, isoelectric point (pI), and GRAVY score (grand average of hydropathicity) using the Expasy ProtParam database (https://web.expasy.org/protparam, accessed on 13 March 2023). Tomato (Solanum lycopersicum), maize (Zea mays), and Arabidopsis thaliana Dof gene family members were also considered in our study.
Identification of domains and phylogenetic analysis of the HuDof genes
Domains of HuDof protein sequences were identified by using the NCBI conserved domain tool (https://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi, accessed on 13 March 2023). We obtained the protein sequences for Dof protein family members of maize (Z. mays – Refgen V4), tomatoes (S. lycopersicum – ITAG4.0), and Arabidopsis (A. thaliana – TAIR-7) from Phytozome-13 (https://phytozome-next.jgi.doe.gov, accessed on 17 March 2023) (Goodstein et al. 2012). Comparative phylogenetic analysis of these plants were carried out by using MEGA X, through maximum likelihood (ML) (Kumar et al. 2018) and the tree was designed through the iTOL database (Letunic and Bork 2019). The boundary was designed through Adobe Illustrator.
Characterisation of gene structure, conserved motif, and sequence logos of HuDof genes
The gene structures of HuDof genes were determined by downloading the pitaya genome from (http://www.pitayagenomic.com, accessed on 25 March 2023). Motif analysis of 26 HuDof genes was carried out by using MEME suite database (https://meme-suite.org/meme/tools/meme, accessed on 13 March 2023), and the motifs’ structures were drawn through TBTools software (Chen et al. 2020). Sequence logos of Dof motifs were created by the WebLogo tool (Crooks et al. 2004).
Chromosomal mapping, duplication, and synteny analysis of HuDof genes on the pitaya chromosomes
Mapping of all 26 HuDof genes on chromosomes was caried out by downloading the pitaya genome (http://www.pitayagenomic.com, accessed on 13 March 2023) (Chen et al. 2022a). The length of the chromosomes was identified through TBTools by using the ‘FASTA stats’ feature (Chen et al. 2020). Its ideogram was generated through PhenoGram (http://visualization.ritchielab.org/phenograms/plot, accessed on 19 March 2023). The output image from this ideogram was used to pinpoint the mapping of all 26 HuDof genes on the chromosomes. Duplication analysis of HuDof genes was carried out through the multiple collinearity scan toolbox (MCScanX) (Chen et al. 2020), with an e-value threshold of 1 × 10−5. Sequences were subjected to one-step MCScanX in the TBTools software (Chen et al. 2020) to create the collinearity data, in order to determine the syntenic connection between the Dof genes of Hylocereus undatus, O. sativa, S. lycopersicum, and A. thaliana. The identified HuDof gene pairs were visualised through CIRCUS analysis.
Sub-cellular localisation, selection pressure analysis, and orthologous gene clustering of HuDof genes
The localisation of HuDof genes was predicted by using Wolf PSORT (https://www.genscript.com/wolf-psort.html, accessed on 25 March 2023) and CELLO (ver. 2.5, http://cello.life.nctu.edu.tw, accessed on 25 March 2023). For selection pressure analysis, stop codons (UAA, UAG, UGA) were removed from HuDof protein sequences and then subjected to analysis in Selecton Server (http://selecton.tau.ac.il/index.html, accessed on 26 March 2023). Using OrthoVenn2, (https://orthovenn2.bioinfotoolkits.net/home, accessed on 28 March 2023) the orthologous genes in H. undatus, Z. mays, S. lycopersicum, and A. thaliana have been detected.
Correlation networks of HuDof genes
We have identified the interactions of all 26 HuDof genes by obtaining data from the pitaya database (http://www.pitayagenomic.com/coexpression.php, accessed on 27 March 2023) (Chen et al. 2022a). Interacting genes were visualised through the Cytoscape program (Shannon et al. 2003).
Identification of promoter and cis-acting elements in HuDof promoter regions
We extracted the data of promoter regions from the pitaya genomic file (2000 bp upstream of the start codon) through TBTool (Chen et al. 2020). For analysis of cis-elements in promoter regions, we used the PlantCARE database (http://bioinformatics.psb.ugent.be/webtools/plantcare/html, accessed on 28 March 2023).
HuDof gene expression profiling by RNA-seq
The expression data of the HuDof genes were retrieved from pitaya database (http://www.pitayagenomic.com, accessed on 26 March 2023) and the circular heat map was constructed through TBTools (Chen et al. 2020). All 26 HuDof genes’ expression levels were evaluated in various plant tissues representing five flower phases (F1, F2, F3, F4, F5), four flowering bud stages (FB1, FB2, FB3, FB4), three pericarp phases (PeriC-45d, PeriC-65d, and PeriC-85d) and the main phases of fruit pulp (Pulp-29d, Pulp-35d, Pulp-49d).
Plant materials and stress treatments
The seeds of pitaya (Selenicereus undatus L.) were collected from Hainan-Shengda Modern Agriculture Development Co., Qionghai, Hainan, China, and grown in a controlled environment at the Nanfan Research Institute in Yazhou Bay Seed Science and Technology City, Hainan University, China. The seeds were sown in trays filled with peat-moss soil and kept in a growth room with a 16-h day (24°C) and an 8-h night (70% relative humidity). After 40 days, seedlings were chosen for heavy metal treatments, coinciding with the onset of stem differentiation into cladodes. Separate experiments were conducted to get the optimised values for each treatment including vanadium (35 mg L−1), 30% 5-polyethylene glycol (PEG-6000), melatonin (100 μM), and 42°C heat by following the protocol (Zaman et al. 2023b).
RNA isolation and RT-qPCR evaluation
Total RNA was extracted from diverse plant tissues, such as stems at different stages (1-month, 1-year, and 2-year), flower buds, pericarp, and pulp by using the RNAprep Pure Plant Kit (TIANGEN, Beijing, China). The NanoDrop 2000C spectrophotometer measured RNA concentrations, and DNase I was applied to eliminate genomic DNA. Subsequently, 1 g of RNA from each sample was reverse transcribed into complementary DNA (cDNA) using the QuantiTect Reverse Transcription Kit. Quantitative reverse transcription PCR (RT-qPCR) was employed to analyse gene expression, with actin as the internal control. Three biological and three technical replications were conducted for each sample. The double-delta CT method was utilised for data analysis, and the SYBER Green Master Mix, along with the LightCycler 480 real-time PCR system, were used in the experiment (Zaman et al. 2023a).
Dof gene family expression studies in response to exogenous melatonin and several stresses
We utilised the FPKM values (fragments per kilobase of transcript per million mapped reads) for the 26 HuDof genes to analyse their expression profiles under both single and multifactorial stress combinations. Plant samples from eight treatments and two replications of single and multifactorial stresses, as described in a previous study (Zaman et al. 2023c), were collected, quickly frozen in liquid nitrogen, and stored at −80°C. The cDNA synthesis process adhered to established protocols from an earlier study (Zaman et al. 2023c). Using RNA-seq data, eight genes were chosen to assess their expression patterns in pitaya tissues across all treatments. To confirm Dof gene expression, RT-qPCR was conducted with primers listed in Supplementary file 1, following methods outlined in a previous study (Zaman et al. 2023c).
Results
Genome-wide identification and analysis of Dof genes in pitaya genome
HuDof-1 to HuDof-26 are the 26 gene family members that were identified by study in the pitaya genome. Physio-chemical characteristics of 26 gene family members were measured, including number of amino acids, molecular weight (kDa), isoelectric point (pI), grand average of hydropathicity (GRAVY), each gene’s chromosomal number, position on the chromosome, coding sequence (CDS) length, protein length, and protein molecular weight. Protein length ranged from 48 amino acids (HU01G02303.1) to 366 amino acids (HU03G01783.1), and their molecular weights ranged from 5 to 40 kDa. Due to the fact that all 26 pitaya HuDof proteins had typical GRAVY values that ranged from −1.586 (HU07G00122.1) to −0.536 (HU02G01516.1), hydrophilicity was predicted for all 26 of them (Table 1). pI net electrical charge at a specific pH value ranged from 9.7 (HU03G01783.1) to 10.5 (HU10G00371.1), with an average value of 9.95 for all HuDof proteins (Supplementary file S1).
| Transcript ID | Renamed ID | Chr. No. | Start position | End position | Protein length | CDS | Molecular weight | pI | GRAVY | |
|---|---|---|---|---|---|---|---|---|---|---|
| HU03G01783.1 | HuDof-1 | 3 | 138123983 | 138125080 | 366 | 1098 | 40 081.22 | 9.17 | −0.781 | |
| HU04G01455.1 | HuDof-2 | 4 | 136752354 | 136752530 | 59 | 177 | 6921.06 | 10.13 | −1.039 | |
| HU01G02303.1 | HuDof-3 | 1 | 19088385 | 19088528 | 48 | 144 | 5658.48 | 9.87 | −0.863 | |
| HU05G01863.1 | HuDof-4 | 5 | 21356210 | 21356416 | 69 | 207 | 7958.24 | 10.25 | −0.862 | |
| HU02G01516.1 | HuDof-5 | 2 | 8900507 | 8900761 | 85 | 255 | 9500.00 | 10.00 | −0.536 | |
| HU01G00880.1 | HuDof-6 | 1 | 60977631 | 60977819 | 63 | 189 | 7328.34 | 9.61 | −1.156 | |
| HU05G02152.1 | HuDof-7 | 5 | 28170303 | 28170461 | 53 | 159 | 6174.09 | 10.07 | −0.919 | |
| HU03G00898.1 | HuDof-8 | 3 | 39827266 | 39827517 | 84 | 252 | 9559.79 | 10.14 | −0.977 | |
| HU01G02405.1 | HuDof-9 | 1 | 122649095 | 122649295 | 67 | 201 | 7779.66 | 9.85 | −1.399 | |
| HU05G00831.1 | HuDof-10 | 5 | 122589317 | 122589514 | 66 | 198 | 7643.57 | 9.6 | −1.056 | |
| HU06G00816.1 | HuDof-11 | 6 | 887677 | 887829 | 51 | 153 | 5903.66 | 9.4 | −0.822 | |
| HU04G02102.1 | HuDof-12 | 4 | 106160814 | 106161074 | 87 | 261 | 10 007.28 | 9.25 | −1.322 | |
| HU06G00823.1 | HuDof-13 | 6 | 105595524 | 105595685 | 54 | 162 | 6306.27 | 10 | −1.011 | |
| HU03G02125.1 | HuDof-14 | 3 | 105185091 | 105185270 | 60 | 180 | 6982.05 | 10.32 | −1.340 | |
| HU05G01867.1 | HuDof-15 | 5 | 4080278 | 4080499 | 60 | 222 | 8617.99 | 10.22 | −0.845 | |
| HU07G00122.1 | HuDof-16 | 7 | 90920517 | 90920750 | 78 | 234 | 9185.44 | 10.31 | −1.586 | |
| HU05G00013.1 | HuDof-17 | 5 | 10068512 | 10068673 | 54 | 162 | 6391.37 | 10 | −1.1 | |
| HU02G00342.1 | HuDof-18 | 2 | 9983757 | 9983912 | 52 | 156 | 6103.91 | 9.6 | −1.012 | |
| HU08G02178.1 | HuDof-19 | 8 | 1614759 | 1614932 | 58 | 174 | 6806.81 | 10.21 | −1.226 | |
| HU08G02142.1 | HuDof-20 | 8 | 83747840 | 83747998 | 53 | 159 | 6186.05 | 9.89 | −0.921 | |
| HU10G01728.1 | HuDof-21 | 10 | 93696298 | 93696459 | 54 | 162 | 6331.2 | 10.14 | −1.076 | |
| HU07G00919.1 | HuDof-22 | 7 | 1268912 | 1269073 | 54 | 162 | 6273.13 | 9.98 | −1.011 | |
| HU04G00361.1 | HuDof-23 | 4 | 18049551 | 18049700 | 50 | 150 | 5744.63 | 9.8 | −0.85 | |
| HU08G02181.1 | HuDof-24 | 8 | 4536199 | 4536426 | 76 | 228 | 8821.97 | 10.1 | −1.309 | |
| HU11G01640.1 | HuDof-25 | 11 | 116532138 | 116532353 | 72 | 216 | 8321.4 | 10.37 | −1.117 | |
| HU10G00371.1 | HuDof-26 | 10 | 28618146 | 28618367 | 74 | 222 | 8798.03 | 10.54 | −1.076 |
Identification of domains
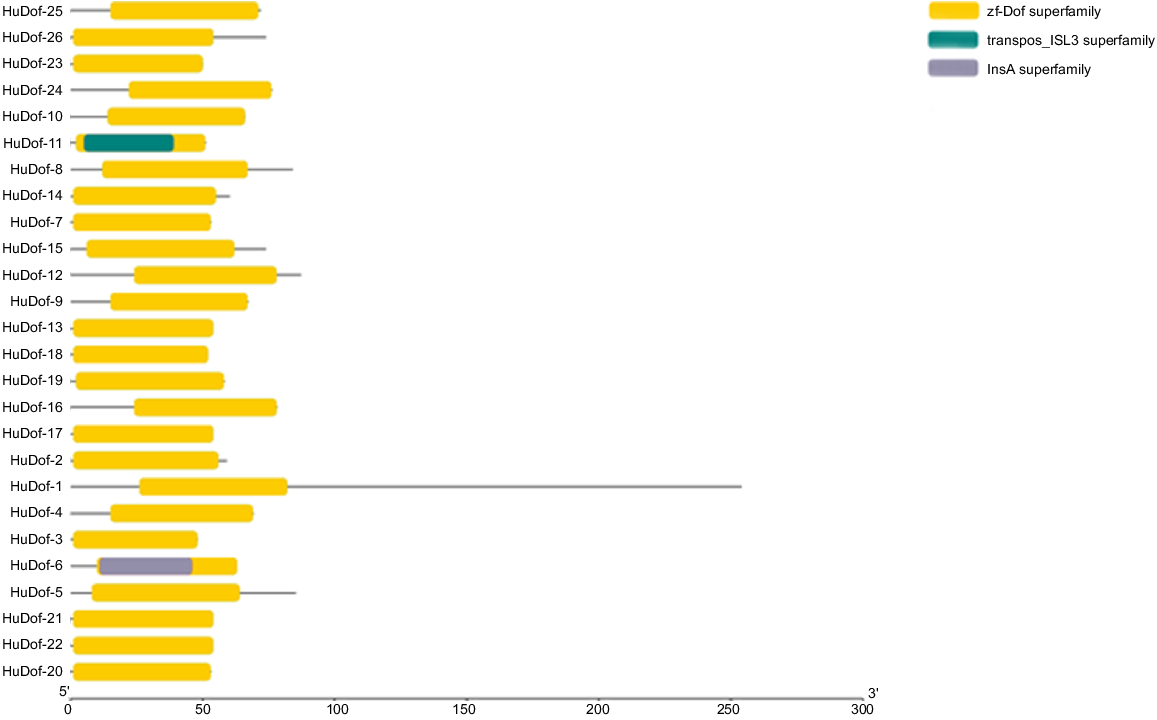
The analysis of domain for all 26 HuDof proteins was retrieved by using the NCBI domain database. The acquired information and TBTools were used to illustrate the domain structure. This domain-based investigation found that the Dof domains were included in all 26 proteins selected for sequencing (Fig. 1).
Motif analysis, gene structure, and phylogenetic tree of HuDof genes
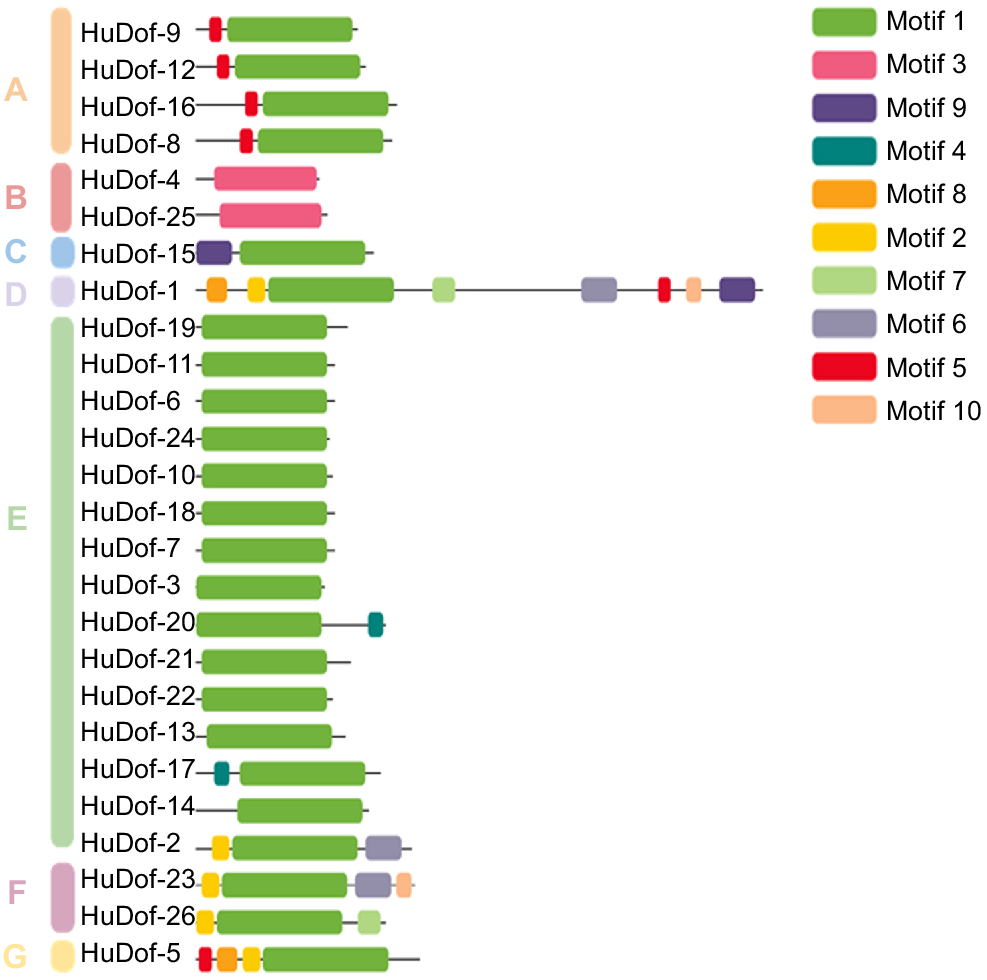
Each Dof protein was examined for homologous motifs using the MEME program. There were ten recognised conserved motifs. Highly conserved domains were present in the C-terminus regions of the HuDof proteins. The majority of the Dof proteins had motifs that showed similarities within the group. Our understanding of the intimate evolutionary ties among the same protein group was also aided by these indications. We clustered them into seven subgroups with appropriate motif sequences. Group A contained four genes, Group B and F contained two genes, Group C, D, and G contained one gene, and Group E contained 15 genes and share similar motif pattern witin the group of HuDof family (Fig. 2).
HuDof-1 to HuDof-26 were analysed for homologous motifs using the MEME program. Motif 1 was found in almost all the HuDof genes. Motif 3, 10, 6, 4, and 9 were present in two different types of genes, and Motif 5 was present in five genes.
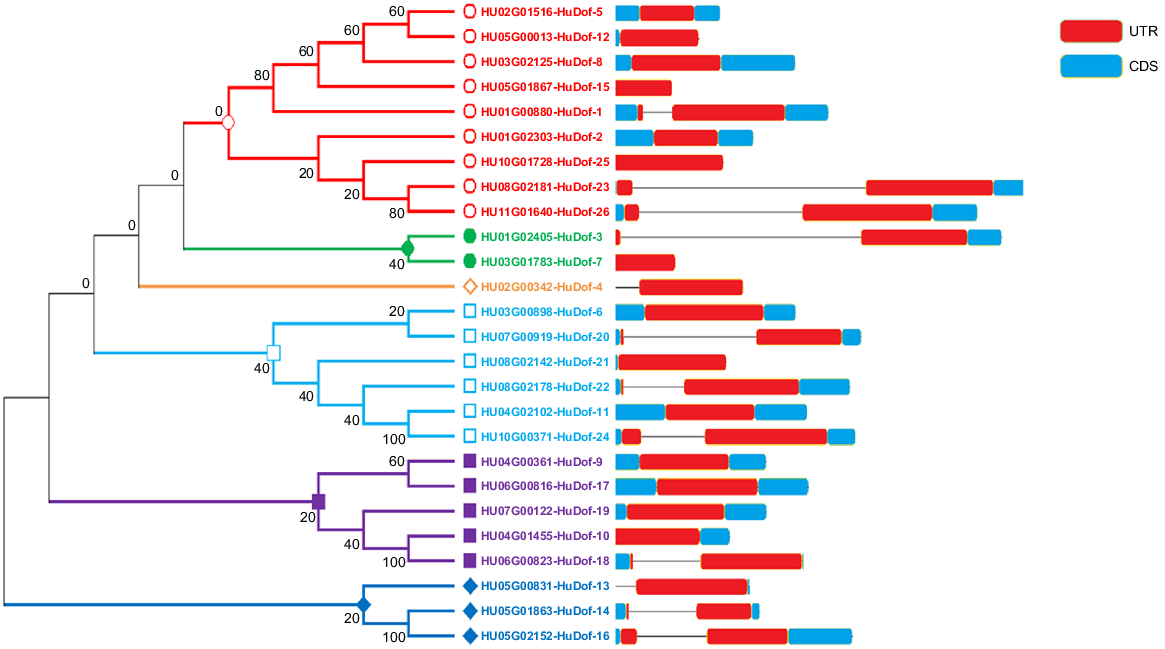
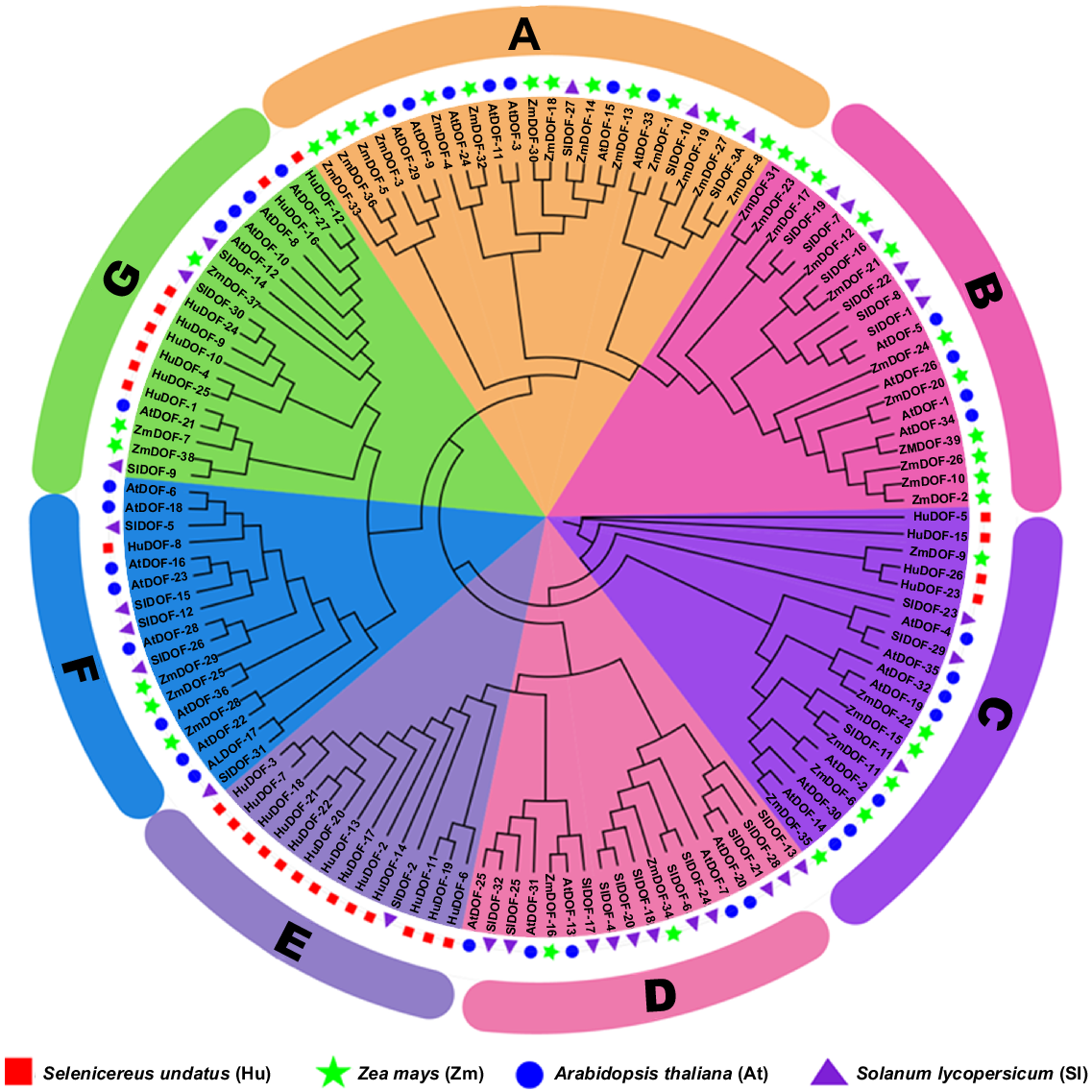
We obtained the Dof proteins from the genomes of each species using the Phytozome database, along with their respective Dof protein sequences from pitaya, maize, tomato, and Arabidopsis (Supplementary file S2). A total of 115 protein sequences were retrieved, including 26 HuDof from pitaya, 37 OsDof from rice, 26 ZmDof from Z. mays, and 26 AtDof from A. thaliana. These were used to construct a phylogenetic tree. The Dof genes were clustered into seven subclasses in the phylogenetic tree on the basis of conserved motifs and their phylogenetic tree (Fig. 3: subfamily A, orange; B, pink; C purple; D, light pink; E, lilac; F, blue; and G, green). Among all, Group E contained the 15 most highly conserved genes and shared similar motif pattern within the gene family, Group C contained 4 genes, Group F contained 1 gene, Group G contained 8 genes.
Comparative phylogenetic analysis of HuDof family proteins of four species, with a bootstrap value of 1000 repetitions. Stars show Zea mays, triangles shows Solanum lycopersicum, circles show Arabidopsis thaliana and squares Selenicereus undatus. Data alignment was carried out using MEGA X software, and the visualisation was crafted with iTool software and finalised with Adobe Illustrator.
It was possible to identify the gene structures by aligning the intron and exon structures of each HuDof gene with a rectangular phylogenetic tree (Fig. 4). Exons and introns can be found in the majority of HuDof gene sequences.
Distribution of HuDof genes amongst pitaya chromosomes, duplication, and synteny analysis
All Dof genes were mapped on specific pitaya chromosomes and illustrated through PhenoGram plot software (Fig. 5). The HuDof genes were irregularly distributed across 11 chromosomes. Chromosome 5 (Chr 5) contained five HuDof genes. There are three HuDof genes in Chr 1, 3, 4, and 8. Two genes on Chr 2, 6, 7, and 10. Chr 11 contains only one gene. Their classification was based on chromosome length and common ancestors in the rectangular phylogenetic tree. The pitaya genome’s chromosome length and each gene’s location are presented in Supplementary file S3.
Dof genes were distributed throughout 11 pitaya chromosomes. The gene names are written in black. Dof genes were divided into six groups on the basis of their domains.

The 26 HuDof genes exhibited an uneven distribution across pitaya’s 11 chromosomes, with five genes on Chr 5, two gene family members on Chr 2, 6, 7, and 10, three on Chr 1, 3, 4, and 8, and one gene on Chr 11. The pitaya genome in the TBTool was examined using the MCScanX software to see if gene duplication events occurred.
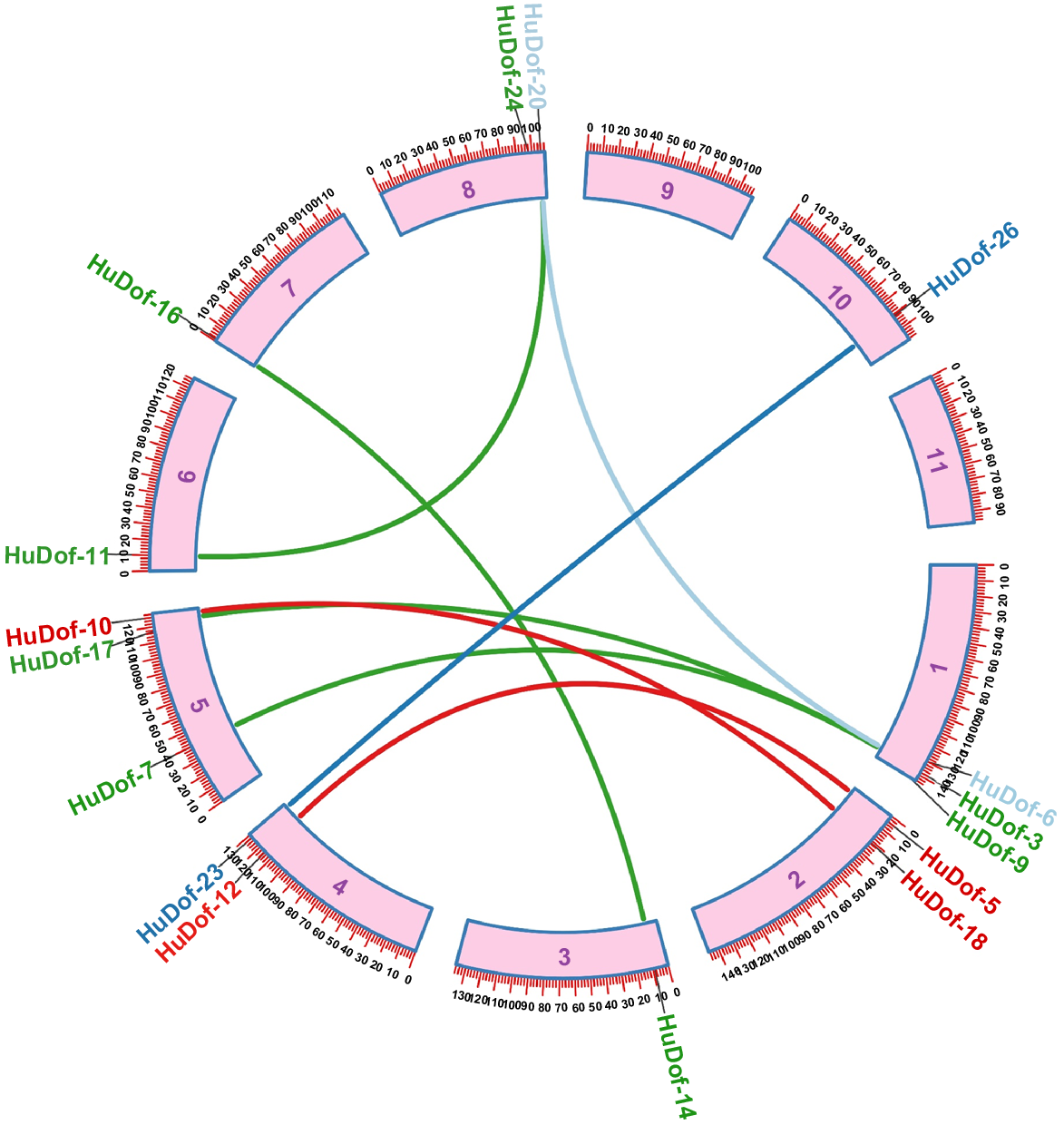
To understand divergence and duplication events among the HuDof gene family members, Ka/Ks ratios for gene pairs were calculated. Divergence time of the gene pairs was computed using the synonymous mutation rate of substitution per synonymous site per year (T = Ks/2x), which was set at x = 6.5 × 10−9 (Zaman et al. 2023b). This helps reveal the gene duplication timing in the HuDof family. Our study identified three pairs of duplication events in HuDofs, which were diverged by segmental duplications in this gene family (Table 2). A circos plot visually displays chromosome connections among HuDof proteins (Fig. 6).
| Gene pair | Ka | Ks | Ka/Ks | Time (mya) | Event | |
|---|---|---|---|---|---|---|
| HuDof3–HuDof7 | 0.51909273 | 1.02691984 | 0.505485 | 0.15 | Segmental duplication | |
| HuDof20–HuDof6 | 0.6731827 | 2.2227868 | 0.302855 | 0.17 | Segmental duplication | |
| HuDof26–HuDof23 | 0.71575747 | 3.26592734 | 0.219159 | 0.25 | Segmental duplication |
Pitaya has duplicated pairs of the HuDof genes, as seen in this circos plot. The genes that have undergone gene duplication events are highlighted in various colours. All pairs show segmental duplications including HuDof23–HuDof26, HuDof3–HuDof7, and HuDof6–HuDof20. However, the rest of the pairs shows duplications, but time of divergence was not predictable. Purple digits indicate Chr numbers.
The nature of selection pressure is identified by using the Ka/Ks ratio. A Ka/Ks value below 1 indicates purifying selection, equal to 1 shows neutral selection and exceeding 1 shows positive selection. The amino acids listed in the positive and purifying selection outcomes are presented using a colour scale (Fig. 7), illustrating that purifying selection has been applied to all proteins.
Purifying selection of HuDof genes. Dark red highlights indicate purifying or negative selection on codons, grey and white highlights indicate neutral selection. Yellow and orange highlights indicate positive selection.

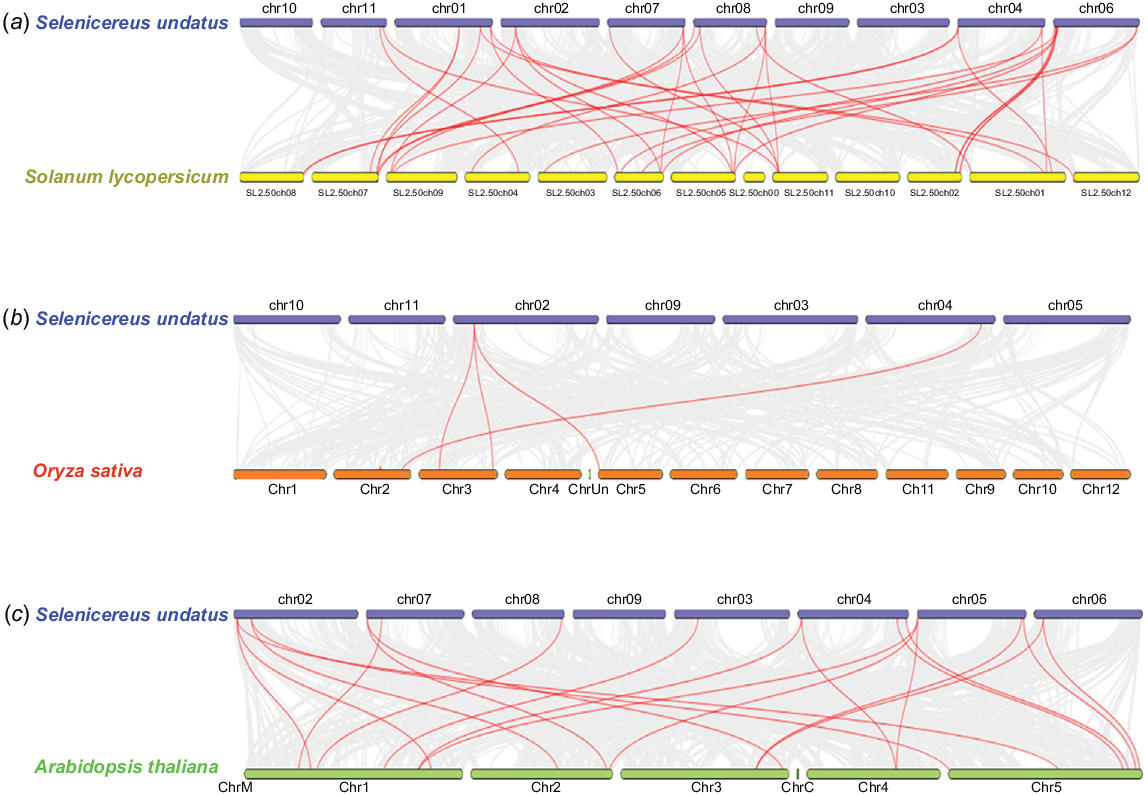
Synteny in S. undatus was assessed in relation to S. lycopersicum, O. sativa, and A. thaliana. Several conserved regions/genes were revealed on a number of chromosomes of the pitaya plant (Fig. 8). The evolutionary relationship of pitaya genes HuDof-6, HuDof-5, HuDof-14, HuDof-12, HuDof-17, HuDof-11, HuDof-16, HuDof-20, HuDof-26, and HuDof-25 was witnessed in relation to Arabidopsis genes AtDof-4, AtDof-10, AtDof-28, AtDof-30, and AtDof-31, respectively.
Visualisation of the syntenic analysis. Synteny analysis of S. undatus with (a) S. lycopersicum (b) O. sativa, and (c) A. thaliana were identified by applying TBtools program to do an MCScanX examination. Grey lines that link the genomes indicate synteny blocks, while red lines indicate the synteny between the genes inside these genomes specifically.
Sub-cellular localisation, selection pressure, and orthologous gene clustering of HuDof genes
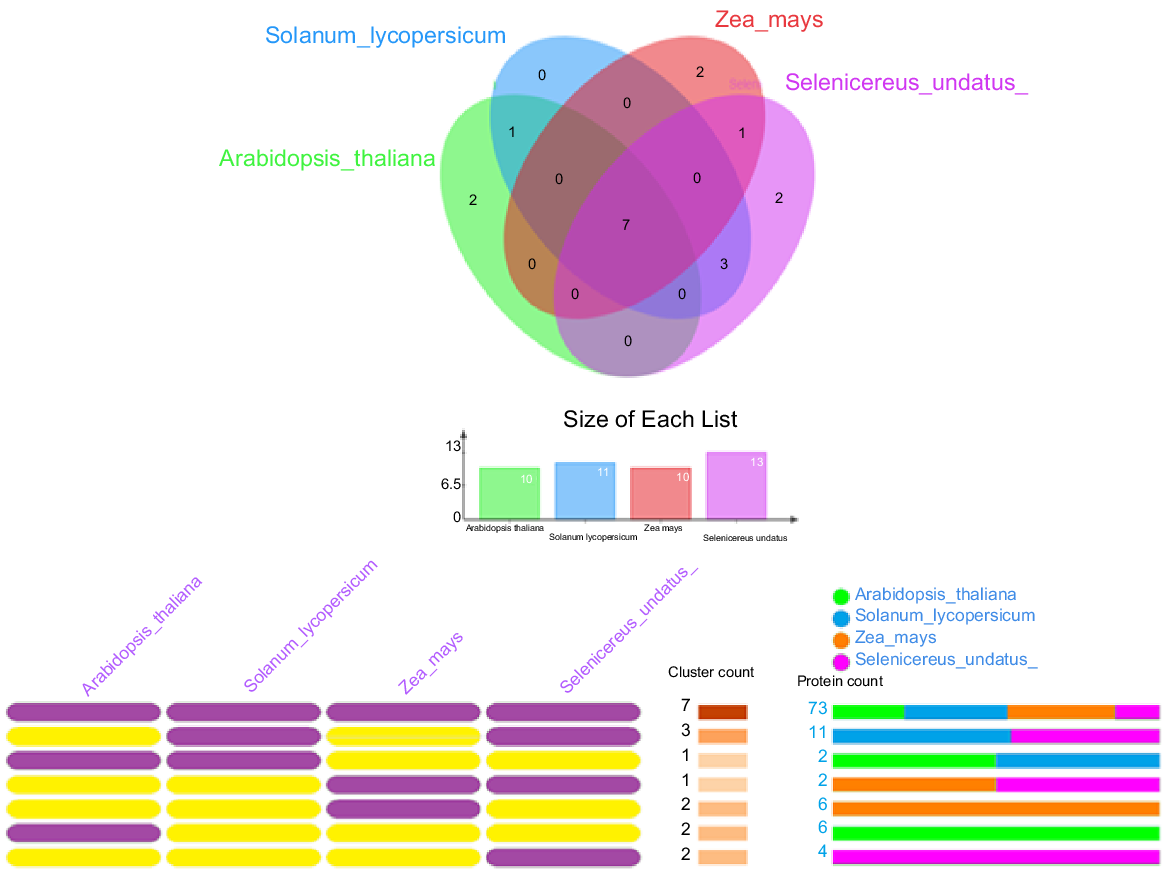
The OrthoVenn2 database was used to study orthologous relationships among Dof genes in four plant species: A. thaliana, Z. mays, S. undatus, and S. lycopersicum. Seventy-three Dof proteins were clustered into seven orthologous groups (Fig. 9).
Orthologous Dof proteins clustering analysis. Clusters of the orthologous protein between the Dof gene family of Z. mays, S. undatus, S. lycopersicum, and A. thaliana have been discovered and visualised by utilising the OrthoVenn2 online tool. For analysis, 1e−10 was used as the e-value cut-off.
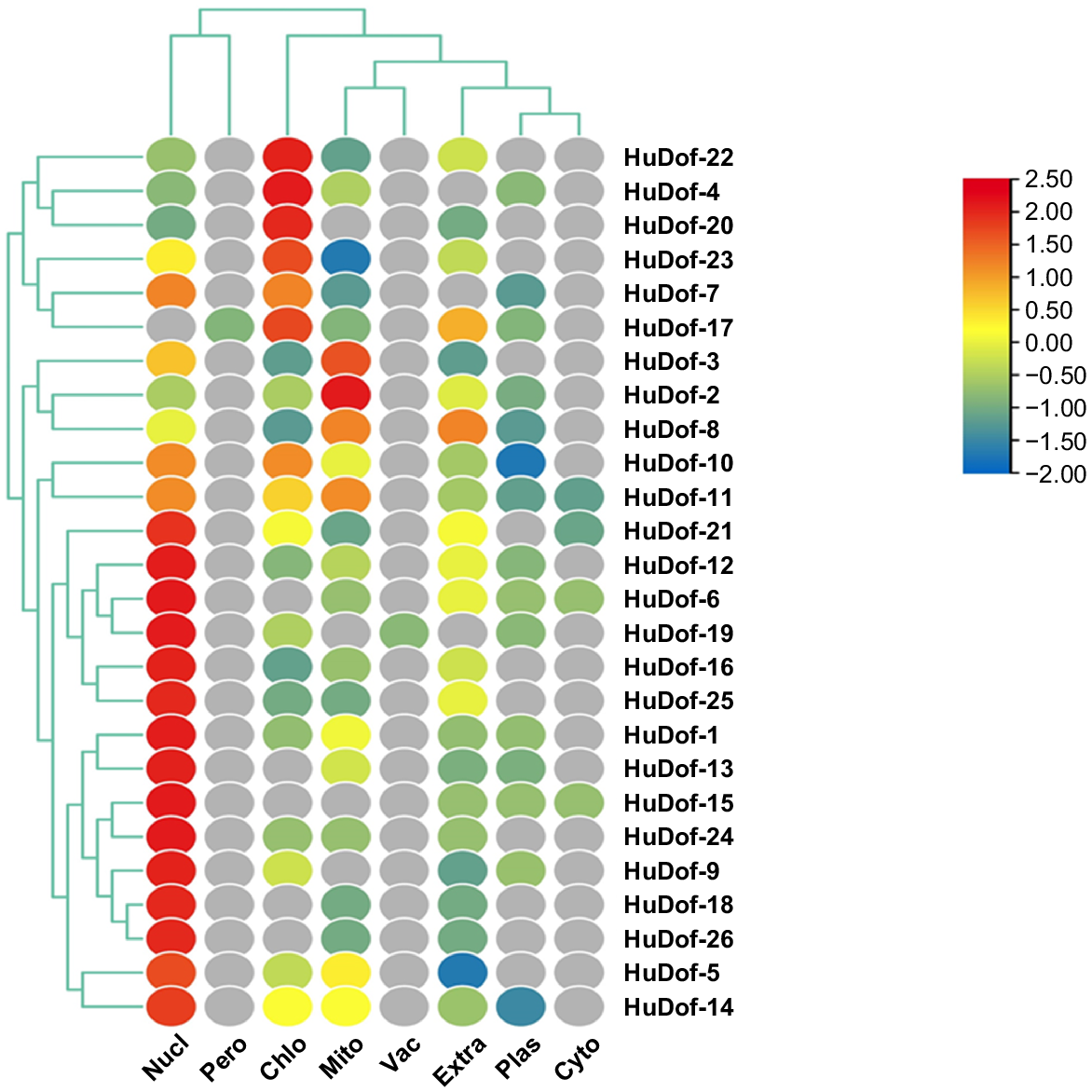
We predicted the role of HuDof genes in several cell organelles, such as the mitochondria, chloroplasts, peroxisomes, and nuclei, using gene localisation (Fig. 10). For example, HuDof-22 was highly localised in chloroplasts, with intermediate levels in the nucleus and lower in the mitochondria. HuDof-4 was also highly localised in chloroplasts, and observed at intermediate levels in nucleus, plastid and mitochondria. Thus, most of the genes were engaged in the nucleus and chloroplasts.
A heat map visually representing HuDof genes’ sub-cellular localisation using a colour scheme (red for high, yellow for intermediate, blue for low protein numbers) in various cellular compartments; i.e. vacuole (Vac), cytoplasm (Cyto), mitochondria (Mito), nucleus (Nucl), plastid (Plas) and peroxisome (Pero). Extra indicates that these genes are localised other than the given organelles in the figure.
HuDof protein network analysis
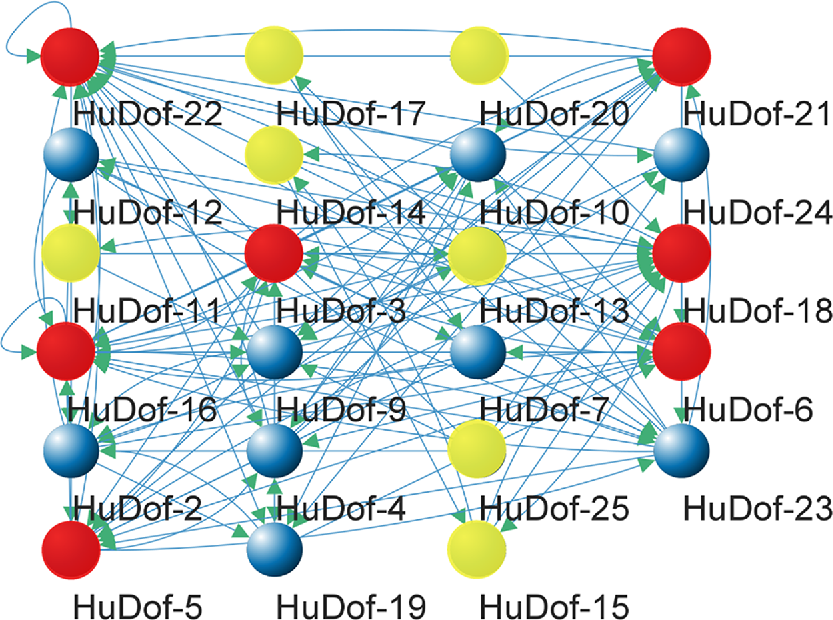
The correlation network of 26 HuDof genes revealed strong interactions among them. HuDof-2, HuDof-5, HuDof-11, HuDof-14, HuDof-18, HuDof-21, and HuDof-22 were more interactive, as shown in Fig. 11 in red.
Protein–protein interactions of HuDof family members in the pitaya. Red shows highest levels of interaction, identified as hub genes. Blue shows intermediate interaction levels and yellow shows weakest interaction. Blue lines show interaction. Green arrows represent the direction of protein interaction.
Cis-acting elements in the promoter region of HuDof genes
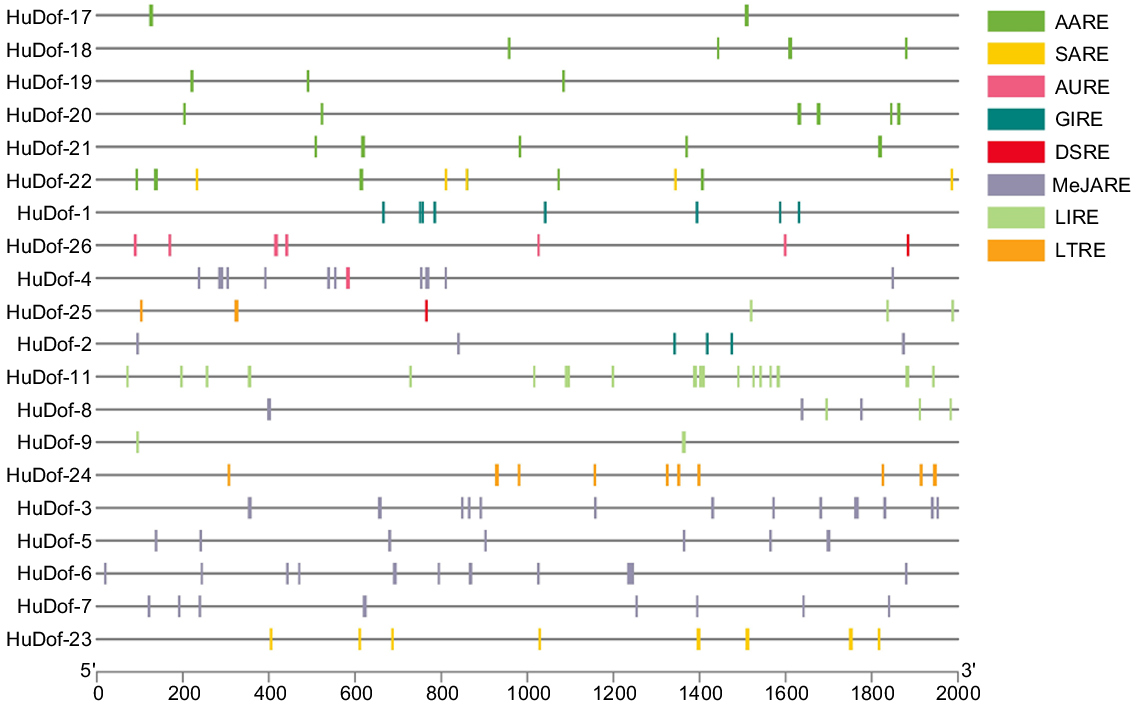
We examined the sequences of the 26 HuDof genes in pitaya (Figs 12 and 13), paying particular attention to the 2000 base pairs upstream of the start codon, to ascertain their biological functions. To locate cis-elements for this investigation, the PlantCARE web tool was used. Nine genes from the HuDof gene family showed elevated expression and a variety of cis-acting regulatory mechanisms (Supplementary file S4). These cis-acting elements fell into eight distinct categories (Fig. 14): AURE (auxin responsive elements), LIRE (light responsive elements), AARE (abscisic acid responsive elements), MejARE (methyl jasmonate responsive elements), DSRE (defence and stress response), GIRE (gibberellin responsive elements), LTRE (low-temperature elements), and SARE (salicylic acid responsive elements).
Twenty-six HuDof genes have cis-acting elements in their promoter regions, which are situated 2000 bp before the start codon. This graphic was built using TBTool.
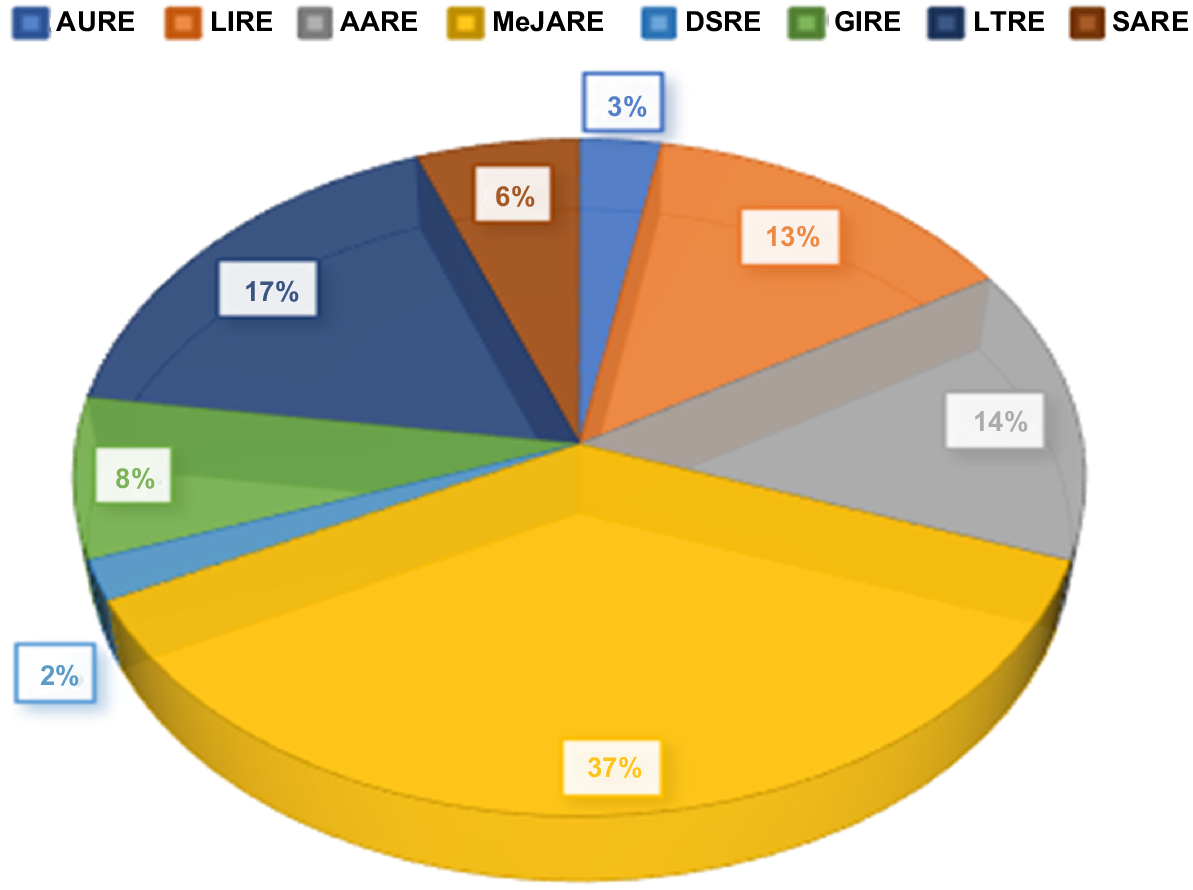
A pie chart showing the different types of elements found in the promoter regions of HuDof genes. These elements are related to various functions, such as responding to light, stress, and hormones like auxin and abscisic acid.
The seven cis-elements linked to the Dof function were examined using the PLACE database. These seven cis-elements were revealed in nearly all the HuDof: the CCAATBOX1, LTRE1HVBLT49, LTRECOREATCOR15, GT1GMSCAM4, MYBCOR, MYCCONSENSUSAT, and DOFCOREZM. DOFCOREZM has also been found in the promoters of genes that are expressed in other parts of the maize plant, such as the roots and leaves, and is involved in various types of functions including development, metabolism, and stress response. MYCCONSENSUSAT plays a role in regulating flowering in soybeans. It is also involved in the promoters of genes that are expressed in other parts of the maize plant, such as the roots and leaves (Supplementary file S5).
Expression analysis of HuDof genes
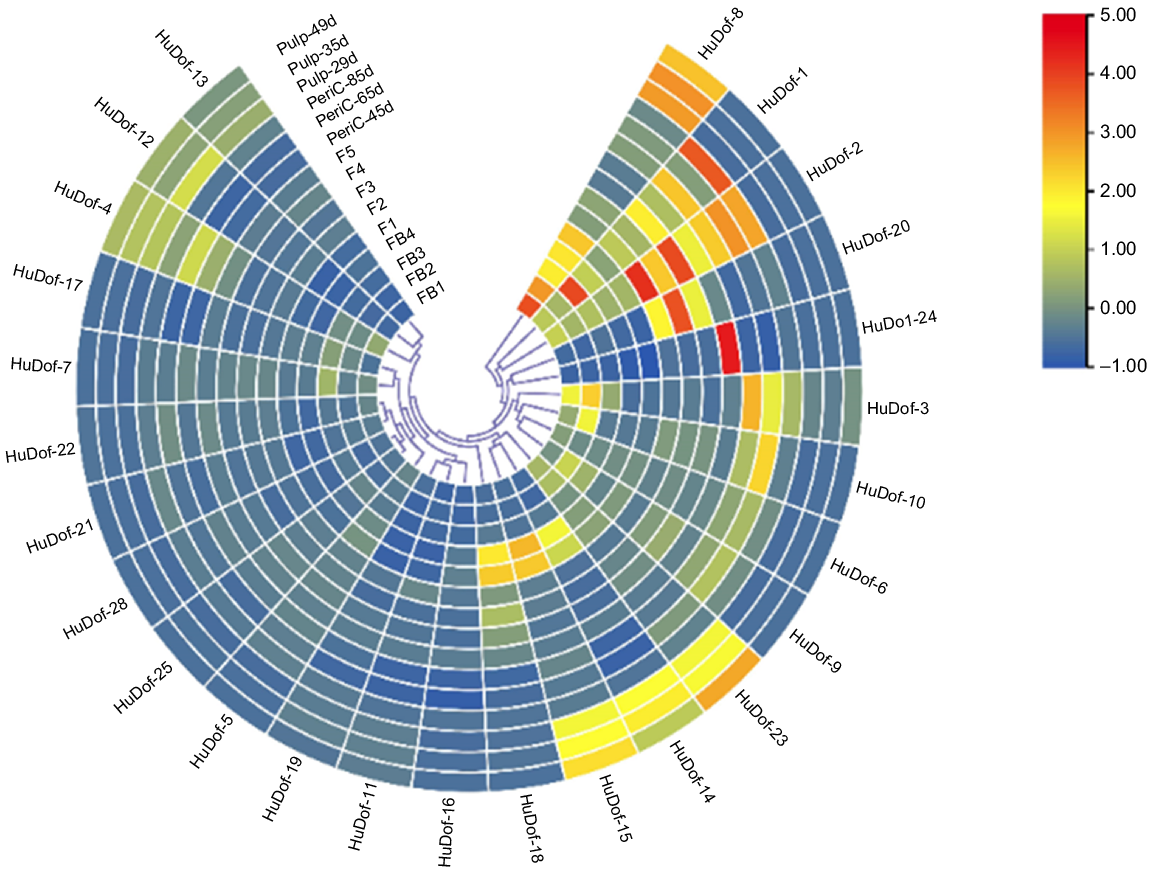
To explore the patterns of HuDof genes, RNA-seq data from 15 tissues was obtained from the pitaya genome database through TBTool. We examined the HuDof gene expression in 15 distinct tissues of the pitaya plant, encompassing pericarp (three stages: 45 days, 65 days, and 85 days), flowers (five stages: F1–F5), flower buds (four stages: FB1–FB4), and fruit pulp (three stages: 29 days, 35 days, and 49 days). To validate the projected gene expressions using the transcriptome data, we conducted RT-qPCR for HuDof-1, HuDof-2, HuDof-3, HuDof-5, HuDof-6, HuDof-8, HuDof-9, HuDof-14, HuDof-23, and HuDof-24 on the basis of high expression in the pitaya plant. Then we designed primers for eight candidate genes. HuDof-1, HuDof-2, and HuDof-8 all had greater levels of expression across the tissues. HuDof-1 exhibited high expression levels across all tissues, including stems, flower buds, and pericarp. However, fruit pulp displayed comparatively lower expression. In contrast, HuDof-2 demonstrated high expression during flowering and pericarp development, with intermediate levels in flower buds. HuDof-8 showed active expression in pulp and flower buds, with weaker expression in flowers.
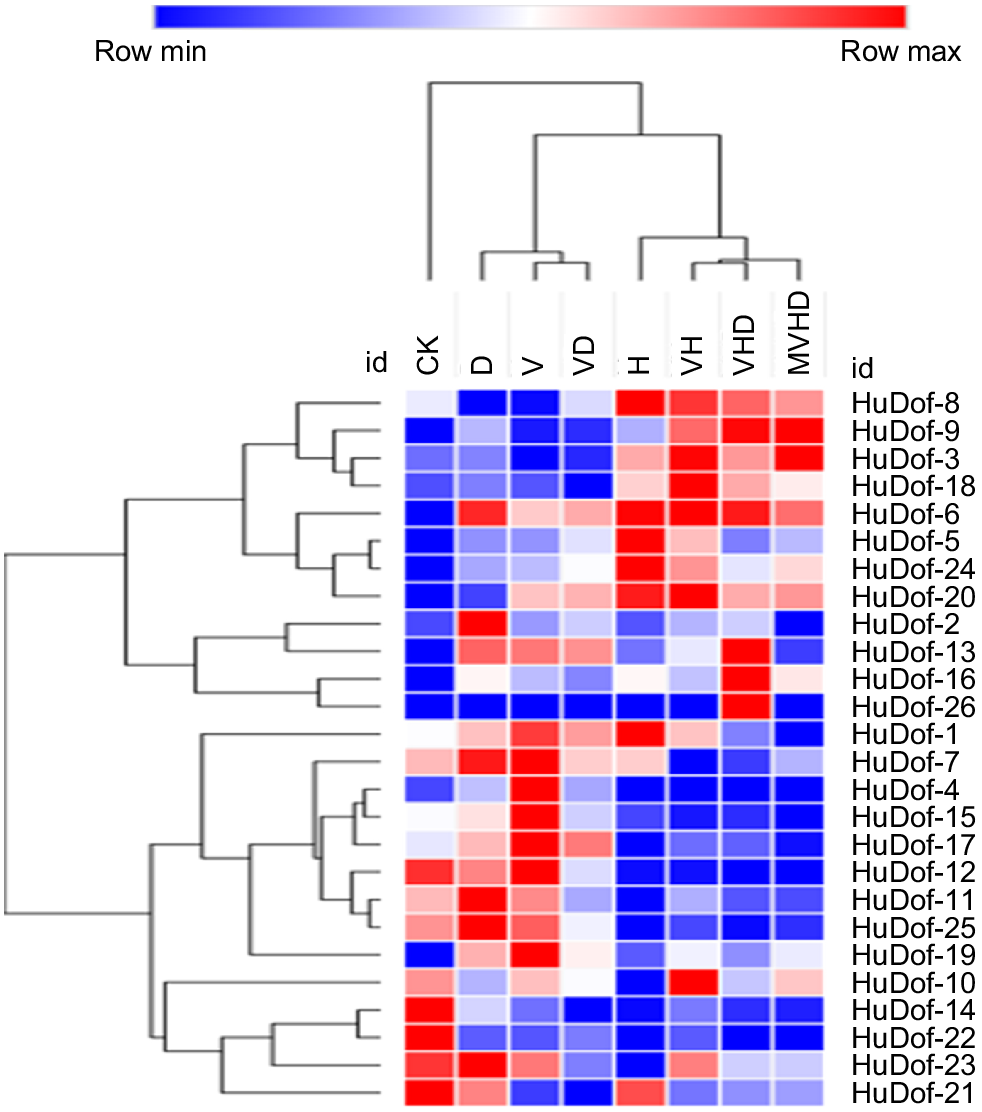
The circular heat map (Fig. 15) shows that how HuDof family genes react to various stress combinations, like drought, heat, vanadium, and others. In the colour bar, red shows highly active genes, with blue indicating less activity. Some genes showed differential expression when the plants were treated with a single stress compared to double abiotic stresses, and even further under multifactorial stresses. Applying melatonin externally boosted the expression of these genes, helping to increase the impact of combined abiotic stresses, which included vanadium, heat, and drought stresses. From the results, HuDof genes play an important role in reducing both single and multiple stresses. Three genes, HuDof-1, HuDof-2, and HuDof-8, showed higher expression in response to specific stresses (D, H, and V) compared to the control (Figs 16 and 17).
The heat map showing the expression of 26 HuDof genes across various pitaya tissues and developmental stages. The 45 columns represent individual genes, and the 15 rows correspond to different tissues and stages, including flower bud, flower, pericarp, and pulp phases. Colour variations within the heat map signify gene expression levels, transitioning from light blue (low or no expression) to dark blue, and from light yellow to dark red (indicating increased expression).
An examination of the HuDof gene family involved a transcriptome analysis under single and multifactorial stress combinations. The expression levels of each gene were depicted using a colour spectrum from dark blue to dark red, measured in log10-based FPKM. The HuDof genes’ expression profiles were assessed under specific stress conditions: CK (control), D (drought), H (heat), V (vanadium), VD (vanadium + drought), VH (vanadium + heat), VHD (vanadium + heat + drought), and MVHD (melatonin + vanadium + heat + drought).
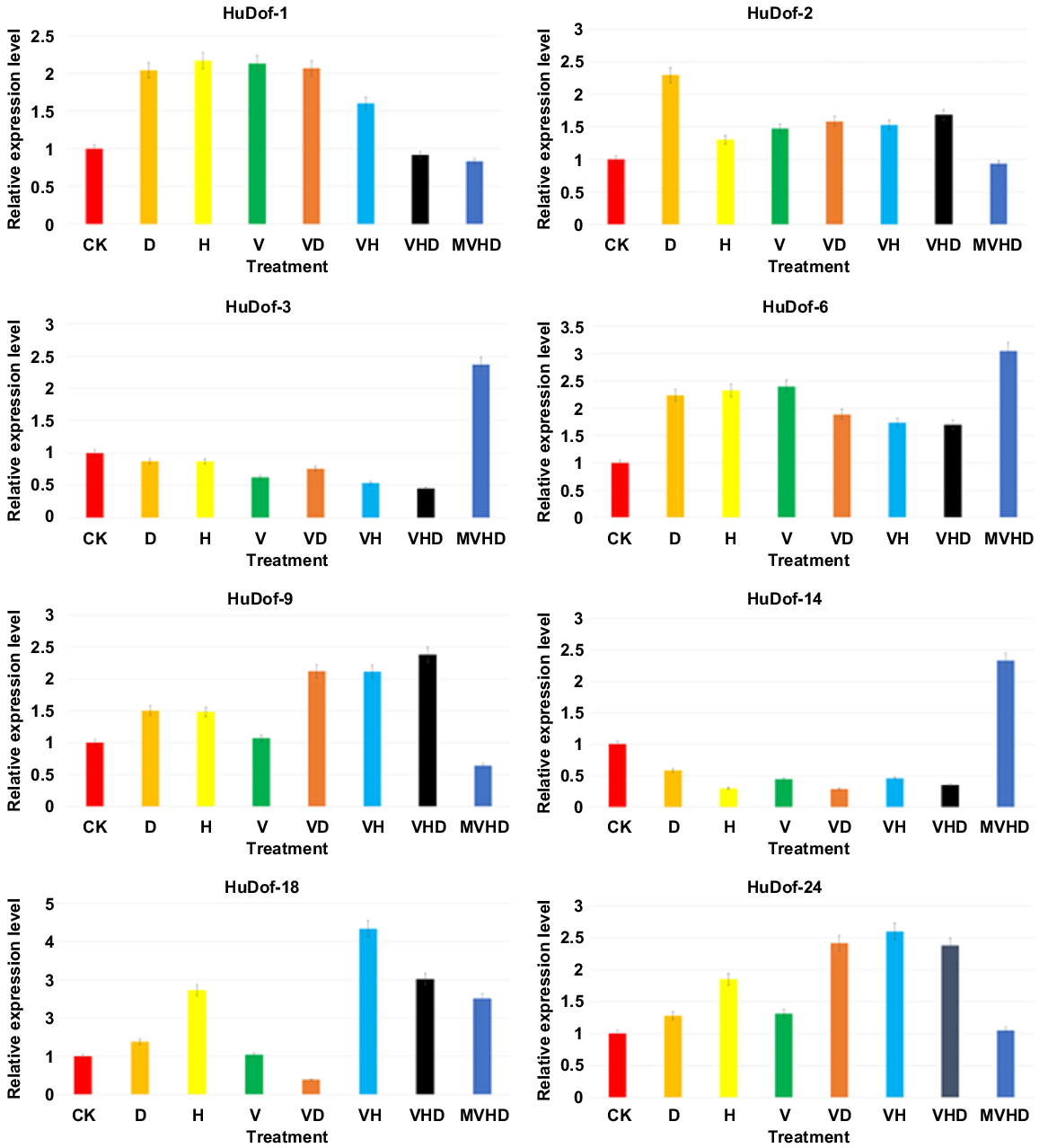
Relative expression analysis of Dof genes was conducted through RT-qPCR. The study involved the validation of the expression of eight HuDof genes in pitaya seedlings subjected to both single and multiple stress conditions. Error bars were included to indicate the standard deviation across replicates. The experimental treatments encompassed eight variations: control (CK), vanadium (V), drought (D), heat (H), vanadium + heat (VH), vanadium + drought (VD), vanadium + heat + drought (VHD), and melatonin + vanadium + heat + drought (MVHD).
Discussion
Pitaya is a tropical fruit of the Cactaceae family, which is evergreen, and consists of cladodes (an altered stem substitutes the leaves to conduct photosynthesis) (Quiroz-González et al. 2018). The pericarp of the fruit is red and the pulp is white in colour, and fruits are nutritious and edible. The fruits include large levels of polyphenols, tannins, betalains, nonbetalainic betalains, and antioxidant substances (Lin et al. 2021). Publication of the genome of white flesh pitaya (Chen et al. 2021) has assisted researchers in identifying candidate genes regulating phenotypic traits. Multiple genes were discovered in pitaya including GRAS (Zaman et al. 2023a), SBP (Khokhar et al. 2023), MATE (Khan et al. 2024), APX (Zaman et al. 2023b), WRKY (Chen et al. 2022b), and HMA (Zaman et al. 2023c). However, an understanding of the Dof gene family in pitaya has remained unavailable. A number of important findings emerged from this study, including the identification of HuDof genes, the physicochemical characteristics of each gene’s CDS sequence, protein length and molecular weight, start and end points on each chromosome, pI, GRAVY, and the predicted sub-cellular localisation of each HuDof family member protein. The Dof transcription factor was present in several plant genomes such as A. thaliana (36 AtDofs) (Yu et al. 2001), rice (30 OsDofs) (Yu et al. 2001), and tomato (34 SlDofs) (The Tomato Genome Consortium 2012). Genome size is 900 Mb in tobacco, (Sierro and Ivanov 2020), 466 Mb in rice, (Vij et al. 2006), 1.41 Gb in pitaya (Chen et al. 2021), and 125 Mb in A. thaliana (Schmuths et al. 2004), suggesting that genome size has no relation to the quantity of Dof proteins.
We have discovered and named the 26 HuDof genes (HuDof 1–26) dispersed across 11 chromosomes in the pitaya genome. We classified the genes into seven subgroups based on conserved motifs, domains structure, and phylogenetic tree, and we drew on the ML approach implemented with MEGA X (Kumar et al. 2018). A similar method was pursued in research on the Dof gene family in watermelon (Zhou et al. 2020).
In the motif analysis, all HuDof proteins across various groups shared similar motifs, except for Motif 1. Notably, each subgroup had its unique and predominantly conserved motifs, with Motif 10 emerging as a secondary conserved domain believed to function as a serine stretch linking the two domains (Ranganathan et al. 1999). Within different subgroups of HuDof genes, the exon–intron structural analysis revealed diverse gene patterns, yet the structural features remained largely similar across these subgroups. HuDofs typically exhibited a low number of introns, ranging from 0 to 2, with most genes possessing only one or none (Fig. 4). These findings suggest that the functions of HuDof genes may diverge significantly among different subgroups.
Gene duplication events, common in the evolution of angiosperms, often include tandem and segmental duplications, which can expand gene families (Wang et al. 2012). Both types of duplications have been reported in related Dof gene families (PtrDofs) (Wang et al. 2017). We used Ka/Ks ratios to understand selection in HuDof genes, estimating gene duplication time. All HuDof proteins are under purifying selection, indicating their conserved functions. This emphasises the significance of preserving specific amino acid positions and protein stability. It reflects the balance between evolutionary pressure and preserving essential protein attributes. Through synteny analysis of S. undatus with S. lycopersicum, O. sativa, and A. thaliana we identified conserved regions on chromosomes of the pitaya plant, helping to elucidate the evolutionary relations and structural characteristics of the HuDof gene family (Yang et al. 2006).
We report a high degree of orthology amonst the species A. thaliana, Z. mays, S. undatus, and S. lycopersicum. This indicates a significant degree of functional redundancy within the Dof gene family, suggesting that Dof proteins in these plants can perform similar functions related to development, growth and stress responses (Fig. 9).
The cis-elements in the promoter region of HuDof genes were analysed using the PLACE database. Three cis-elements (CCAATBOX1, LTRE1HVBLT49, and LTRECOREATCOR15) (Shah et al. 2024) were involved in the metabolic regulation of heat stress also observed in N. tabacum (Fornara et al. 2009). GT1GMSCAM4 helps in the regulation of salinity stress (Senavirathne et al. 2017). However, the DOFCOREZM is involved in plant metabolism and responding to drought conditions (Song et al. 2019). The presence of these stress-responsive cis-elements indicates that HuDof genes have diverse roles in responding to abiotic stresses that affect the growth and productivity of plants (Raza et al. 2022; Khan et al. 2024).
We exposed pitaya seedlings to various single and multifactorial stress conditions, showing that HuDof genes play an important role in reducing both types of stress. Notably, we identified three putative candidate genes (HuDof-1, HuDof-2, and HuDof-8) with potential for enhancing the tolerance in pitaya. Experimental analysis has revealed that pitaya plants activate a variety of genes under the influence of different stresses (Mittler 2006). The response to stresses, faced individually or in combination, can be better understood through analysis of genetic data. In the future, we can create more robust pitaya types by using genetic engineering to comprehend and enhance the role of highly active genes and stress-related components.
Determining how HuDof genes are related to others in the same gene family helps to identify genes that regulate specific plant characteristics. The pathways may be used to identify both positive and negative regulators; then a pitaya plant devoid of transgenes may be produced using the CRISPR technology (Zaman et al. 2023d). In the past, several crops have been enhanced utilising the most recent genome editing method (Khatodia et al. 2016; Haque et al. 2018; Zaman et al. 2019a).
The genome editing method has been used to enhance numerous crop qualities (Zaman et al. 2019b), through study of specific traits. In this study, we identified three putative candidate genes (HuDof-1, HuDof-2, and HuDof-8) using a genome-wide study and RNA-sequence data analysis. These genes are responsible for combating both single and multiple stresses. Our findings related to Dof genes in pitaya growth provide a theoretical framework to enhance the pitaya breeding program.
Conclusion
In this analysis, 26 HuDof proteins were identified, and divided into seven subgroups. The HuDof genes have distinct subgroup-specific gene structures and arrangements of motifs in each subgroup. The HuDof gene family has mostly developed through segmental duplication. HuDof genes also had preferences for expression in distinct tissue and fruit production stages, the majority of which were significantly influenced by the administration of exogenous hormones. Based on our transcriptome data, putative candidate genes such as HuDof-1, HuDof-2, and HuDof-8 have been identified. These genes may have diverse roles in enhancing the tolerance against single, double and multifactorial stresses and can be utilised in traditional and molecular breeding programs aimed at developing pitaya genotypes resilient to climate variations. This finding implies that HuDof genes may be active in a number of metabolic processes throughout the growth and development of pitayas.
Data availability
The data generated and analysed in this study are available in the supplementary materials.
Declaration of funding
This study was funded by the Hainan Province Science and Technology Special Fund (ZDYF2022XDNY190), the Project of Sanya Yazhou Bay Science and Technology City (Grant Number: SCKJ-JYRC-2022-83), Collaborative Innovation Centre of Nanfan and High-Efficiency Tropical Agriculture, Hainan University Funding (XTCX2022NYB09) and the Hainan Provincial Natural Science Foundation of China (421RC486). This project was supported by Researchers Supporting Project number (RSP2024R385), King Saud University, Riyadh, Saudi Arabia.
Author contributions
O.A. and L.U.K. devised the study objective, Q.U.Z. and H.F.W. designed the experiment, O.A. and M.A. analysed the data and interpreted the results. L.U.K. performed the expression analysis for Dof gene family members. Q.U.Z. provided material help to perform the experiment. A.K., M.A., F.M., S.H.S., M.J.A., and Q.U.Z. helped to edit and proofread the manuscript. H.F.W. provided funds and supervised the research.
References
Cai X, Zhang Y, Zhang C, Zhang T, Hu T, Ye J, Zhang J, Wang T, Li H, Ye Z (2013) Genome-wide analysis of plant-specific Dof transcription factor family in tomato. Journal of Integrative Plant Biology 55(6), 552-566.
| Crossref | Google Scholar | PubMed |
Chen Y, Cao J (2015) Comparative analysis of Dof transcription factor family in maize. Plant Molecular Biology Reporter 33, 1245-1258.
| Crossref | Google Scholar |
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, Xia R (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Molecular Plant 13(8), 1194-1202.
| Crossref | Google Scholar | PubMed |
Chen J-Y, Xie F-F, Cui Y-Z, Chen C-B, Lu W-J, Hu X-D, Hua Q-Z, Zhao J, Wu Z-J, Gao D, Zhang Z-K, Jiang W-K, Sun Q-M, Hu G-B, Qin Y-H (2021) A chromosome-scale genome sequence of pitaya (Hylocereus undatus) provides novel insights into the genome evolution and regulation of betalain biosynthesis. Horticulture Research 8, 164.
| Crossref | Google Scholar |
Chen C, Li F, Xie F, Chen J, Hua Q, Chen J, Wu Z, Zhang Z, Zhang R, Zhao J, Hu G, Qin Y (2022a) Pitaya genome and multiomics database (PGMD): a comprehensive and integrative resource of Selenicereus undatus. Genes 13(5), 745.
| Crossref | Google Scholar | PubMed |
Chen C, Xie F, Shah K, Hua Q, Chen J, Zhang Z, Zhao J, Hu G, Qin Y (2022b) Genome-wide identification of WRKY gene family in pitaya reveals the involvement of HmoWRKY42 in betalain biosynthesis. International Journal of Molecular Sciences 23(18), 10568.
| Crossref | Google Scholar | PubMed |
Crooks GE, Hon G, Chandonia J-M, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Research 14(6), 1188-1190.
| Crossref | Google Scholar | PubMed |
Fornara F, Panigrahi KCS, Gissot L, Sauerbrunn N, Rühl M, Jarillo JA, Coupland G (2009) Arabidopsis DOF transcription factors act redundantly to reduce CONSTANS expression and are essential for a photoperiodic flowering response. Developmental Cell 17(1), 75-86.
| Crossref | Google Scholar | PubMed |
Goodstein DM, Shu S, Howson R, Neupane R, Hayes RD, Fazo J, Mitros T, Dirks W, Hellsten U, Putnam N, Rokhsar DS (2012) Phytozome: a comparative platform for green plant genomics. Nucleic Acids Research 40(D1), D1178-D1186.
| Crossref | Google Scholar |
Gualberti G, Papi M, Bellucci L, Ricci I, Bouchez D, Camilleri C, Costantino P, Vittorioso P (2002) Mutations in the Dof zinc finger genes DAG2 and DAG1 influence with opposite effects the germination of Arabidopsis seeds. The Plant Cell 14(6), 1253-1263.
| Crossref | Google Scholar | PubMed |
Gupta S, Malviya N, Kushwaha H, Nasim J, Bisht NC, Singh VK, Yadav D (2015) Insights into structural and functional diversity of Dof (DNA binding with one finger) transcription factor. Planta 241, 549-562.
| Crossref | Google Scholar | PubMed |
Haque E, Taniguchi H, Hassan MM, Bhowmik P, Karim MR, Śmiech M, Zhao K, Rahman M, Islam T (2018) Application of CRISPR/Cas9 genome editing technology for the improvement of crops cultivated in tropical climates: recent progress, prospects, and challenges. Frontiers in Plant Science 9, 617.
| Crossref | Google Scholar | PubMed |
Khan D, Hui L, Khokhar AA, Hussain MA, Lv W, Zaman QU, Wang H-F (2024) Functional characterization of MATE gene family under abiotic stresses and melatonin-mediated tolerance in dragon fruit (Selenicereus undatus L.). Plant Stress 11, 100300.
| Crossref | Google Scholar |
Khatodia S, Bhatotia K, Passricha N, Khurana SMP, Tuteja N (2016) The CRISPR/Cas genome-editing tool: application in improvement of crops. Frontiers in Plant Science 7, 506.
| Crossref | Google Scholar | PubMed |
Khokhar AA, Hui L, Khan D, Hussain MA, Lv W, Zaman QU, Wang H-F (2023) Comprehensive characterization of SBP genes revealed their role under multifactorial stress combinations in dragon fruit (Selenicereus undatus L.). Plant Stress 10, 100294.
| Crossref | Google Scholar |
Konishi M, Yanagisawa S (2007) Sequential activation of two Dof transcription factor gene promoters during vascular development in Arabidopsis thaliana. Plant Physiology and Biochemistry 45(8), 623-629.
| Crossref | Google Scholar | PubMed |
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution 35(6), 1547-1549.
| Crossref | Google Scholar | PubMed |
Kushwaha H, Gupta S, Singh VK, Rastogi S, Yadav D (2011) Genome wide identification of Dof transcription factor gene family in sorghum and its comparative phylogenetic analysis with rice and Arabidopsis. Molecular Biology Reports 38, 5037-5053.
| Crossref | Google Scholar | PubMed |
Letunic I, Bork P (2019) Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Research 47(W1), W256-W259.
| Crossref | Google Scholar | PubMed |
Lijavetzky D, Carbonero P, Vicente-Carbajosa J (2003) Genome-wide comparative phylogenetic analysis of the rice and Arabidopsis Dof gene families. BMC Evolutionary Biology 3(1), 17.
| Crossref | Google Scholar |
Lin X, Gao H, Ding Z, Zhan R, Zhou Z, Ming J (2021) Comparative metabolic profiling in pulp and peel of green and red pitayas (Hylocereus polyrhizus and Hylocereus undatus) reveals potential valorization in the pharmaceutical and food industries. BioMed Research International 2021, 6546170.
| Crossref | Google Scholar |
Liu Y, Liu N, Deng X, Liu D, Li M, Cui D, Hu Y, Yan Y (2020) Genome-wide analysis of wheat DNA-binding with one finger (Dof) transcription factor genes: evolutionary characteristics and diverse abiotic stress responses. BMC Genomics 21(1), 276.
| Crossref | Google Scholar |
Marzábal P, Gas E, Fontanet P, Vicente-Carbajosa J, Torrent M, Ludevid MD (2008) The maize Dof protein PBF activates transcription of γ-zein during maize seed development. Plant Molecular Biology 67, 441-454.
| Crossref | Google Scholar | PubMed |
Mittler R (2006) Abiotic stress, the field environment and stress combination. Trends in Plant Science 11(1), 15-19.
| Crossref | Google Scholar | PubMed |
Moreno-Risueno MÁ, Martínez M, Vicente-Carbajosa J, Carbonero P (2007) The family of DOF transcription factors: from green unicellular algae to vascular plants. Molecular Genetics and Genomics 277, 379-390.
| Crossref | Google Scholar | PubMed |
Negi J, Moriwaki K, Konishi M, Yokoyama R, Nakano T, Kusumi K, Hashimoto-Sugimoto M, Schroeder JI, Nishitani K, Yanagisawa S, Iba K (2013) A Dof transcription factor, SCAP1, is essential for the development of functional stomata in Arabidopsis. Current Biology 23(6), 479-484.
| Crossref | Google Scholar | PubMed |
Noguero M, Atif RM, Ochatt S, Thompson RD (2013) The role of the DNA-binding One Zinc Finger (DOF) transcription factor family in plants. Plant Science 209, 32-45.
| Crossref | Google Scholar | PubMed |
Paśko P, Galanty A, Zagrodzki P, Ku YG, Luksirikul P, Weisz M, Gorinstein S (2021) Bioactivity and cytotoxicity of different species of pitaya fruits – a comparative study with advanced chemometric analysis. Food Bioscience 40, 100888.
| Crossref | Google Scholar |
Quiroz-González B, García-Mateos R, Corrales-García JJE, Colinas-León MT (2018) Pitaya (Stenocereus spp.): an under-utilized fruit. Journal of the Professional Association for Cactus Development 20, 82-100.
| Crossref | Google Scholar |
Ranganathan S, Simpson KJ, Shaw DC, Nicholas KR (1999) The whey acidic protein family: a new signature motif and three-dimensional structure by comparative modeling. Journal of Molecular Graphics and Modelling 17(2), 106-113.
| Crossref | Google Scholar | PubMed |
Raza A, Salehi H, Rahman MA, Zahid Z, Madadkar Haghjou M, Najafi-Kakavand S, Charagh S, Osman HS, Albaqami M, Zhuang Y, Siddique KHM, Zhuang W (2022) Plant hormones and neurotransmitter interactions mediate antioxidant defenses under induced oxidative stress in plants. Frontiers in Plant Science 13, 961872.
| Crossref | Google Scholar |
Ruiz-Domínguez C, Vovides AP, Sosa V (2019) Systematic relevance of pollen morphology in tribe Hylocereeae (Cactaceae). PhytoKeys 128, 121-140.
| Crossref | Google Scholar | PubMed |
Santos LA, de Souza SR, Fernandes MS (2012) OsDof25 expression alters carbon and nitrogen metabolism in Arabidopsis under high N-supply. Plant Biotechnology Reports 6, 327-337.
| Crossref | Google Scholar |
Schmuths H, Meister A, Horres R, Bachmann K (2004) Genome size variation among accessions of Arabidopsis thaliana. Annals of Botany 93(3), 317-321.
| Crossref | Google Scholar | PubMed |
Senavirathne WMA, Jayatilake DV, Herath V, Wickramasinghe HAM (2017) Evaluation of genetic diversity of cis-acting elements of Abscisic acid responsive element binding protein (ABRE-BP) in selected Sri Lankan rice varieties. Tropical Agricultural Research 28, 120-132.
| Crossref | Google Scholar |
Shah Z, Ahmad S, Ullah A, Munir I, Munir U, Khan A, Sultan W (2024) Insilico-based genome-wide identification and structural characterization of heat shock proteins 70 (HSP70) in cowpea (Vigna unguiculata L.). Genetic Resources and Crop Evolution 71, 1053-1065.
| Crossref | Google Scholar |
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Research 13(11), 2498-2504.
| Crossref | Google Scholar | PubMed |
Song Z, Pan F, Lou X, Wang D, Yang C, Zhang B, Zhang H (2019) Genome-wide identification and characterization of Hsp70 gene family in Nicotiana tabacum. Molecular Biology Reports 46(2), 1941-1954.
| Crossref | Google Scholar | PubMed |
Su Y, Liang W, Liu Z, Wang Y, Zhao Y, Ijaz B, Hua J (2017) Overexpression of GhDof1 improved salt and cold tolerance and seed oil content in Gossypium hirsutum. Journal of Plant Physiology 218, 222-234.
| Crossref | Google Scholar | PubMed |
The Tomato Genome Consortium (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature 485(7400), 635-641.
| Crossref | Google Scholar |
Todeschini A-L, Georges A, Veitia RA (2014) Transcription factors: specific DNA binding and specific gene regulation. Trends in Genetics 30(6), 211-219.
| Crossref | Google Scholar | PubMed |
Umemura Y, Ishiduka T, Yamamoto R, Esaka M (2004) The Dof domain, a zinc finger DNA-binding domain conserved only in higher plants, truly functions as a Cys2/Cys2 Zn finger domain. The Plant Journal 37(5), 741-749.
| Crossref | Google Scholar | PubMed |
Venkatesh J, Park SW (2015) Genome-wide analysis and expression profiling of DNA-binding with one zinc finger (Dof) transcription factor family in potato. Plant Physiology and Biochemistry 94, 73-85.
| Crossref | Google Scholar | PubMed |
Vij S, Gupta V, Kumar D, Vydianathan R, Raghuvanshi S, Khurana P, Khurana JP, Tyagi AK (2006) Decoding the rice genome. BioEssays 28(4), 421-432.
| Crossref | Google Scholar | PubMed |
Wang Y, Wang X, Paterson AH (2012) Genome and gene duplications and gene expression divergence: a view from plants. Annals of the New York Academy of Sciences 1256(1), 1-14.
| Crossref | Google Scholar |
Wang H, Zhao S, Gao Y, Yang J (2017) Characterization of Dof transcription factors and their responses to osmotic stress in poplar (Populus trichocarpa). PLoS ONE 12(1), e0170210.
| Crossref | Google Scholar | PubMed |
Yanagisawa S (2002) The Dof family of plant transcription factors. Trends in Plant Science 7(12), 555-560.
| Crossref | Google Scholar | PubMed |
Yanagisawa S, Schmidt RJ (1999) Diversity and similarity among recognition sequences of Dof transcription factors. The Plant Journal 17(2), 209-214.
| Crossref | Google Scholar | PubMed |
Yanagisawa S, Sheen J (1998) Involvement of maize Dof zinc finger proteins in tissue-specific and light-regulated gene expression. The Plant Cell 10(1), 75-89.
| Crossref | Google Scholar | PubMed |
Yang X, Tuskan GA, Cheng (Max) Z-M (2006) Divergence of the Dof gene families in poplar, Arabidopsis, and rice suggests multiple modes of gene evolution after duplication. Plant Physiology 142(3), 820-830.
| Crossref | Google Scholar | PubMed |
Yang J, Yang M-F, Zhang W-P, Chen F, Shen S-H (2011) A putative flowering-time-related Dof transcription factor gene, JcDof3, is controlled by the circadian clock in Jatropha curcas. Plant Science 181(6), 667-674.
| Crossref | Google Scholar | PubMed |
Yu J, Hu S, Wang J, Li S, Wong K-SG, Liu B, Deng Y, Dai L, Zhou Y, Zhang X, et al. (2001) A draft sequence of the rice (Oryza sativa ssp. indica) genome. Chinese Science Bulletin 46, 1937-1942.
| Crossref | Google Scholar |
Zaman QU, Chu W, Hao M, Shi Y, Sun M, Sang S-F, Mei D, Cheng H, Liu J, Li C, Hu Q (2019a) CRISPR/Cas9-mediated multiplex genome editing of JAGGED gene in Brassica napus L. Biomolecules 9(11), 725.
| Crossref | Google Scholar | PubMed |
Zaman QU, Li C, Cheng H, Hu Q (2019b) Genome editing opens a new era of genetic improvement in polyploid crops. The Crop Journal 7(2), 141-150.
| Crossref | Google Scholar |
Zaman QU, Hussain MA, Khan LU, Cui J-P, Hui L, Khan D, Lv W, Wang H-F (2023a) Genome-wide identification and expression pattern of the GRAS gene family in pitaya (Selenicereus undatus L.). Biology 12(1), 11.
| Crossref | Google Scholar | PubMed |
Zaman QU, Hussain MA, Khan LU, Hui L, Khan D, Khokhar AA, Cui J, Raza A, Wang H-F (2023b) Genome-wide identification and expression profiling of APX gene family under multifactorial stress combinations and melatonin-mediated tolerance in pitaya. Scientia Horticulturae 321, 112312.
| Crossref | Google Scholar |
Zaman QU, Khan LU, Hussain MA, Ali A, Hui L, Khokhar AA, Khan D, Wang H-F (2023c) Characterizing the HMA gene family in dragon fruit (Selenicereus undatus L.) and revealing their response to multifactorial stress combinations and melatonin-mediated tolerance. South African Journal of Botany 163, 145-156.
| Crossref | Google Scholar |
Zaman QU, Raza A, Lozano-Juste J, Chao L, Jones MGK, Wang H-F, Varshney RK (2023d) Engineering plants using diverse CRISPR-associated proteins and deregulation of genome-edited crops. Trends in Biotechnology
| Crossref | Google Scholar |
Zhou Y, Cheng Y, Wan C, Li J, Yang Y, Chen J (2020) Genome-wide characterization and expression analysis of the Dof gene family related to abiotic stress in watermelon. PeerJ 8, e8358.
| Crossref | Google Scholar | PubMed |