Genome-wide identification, phylogeny and expression analysis of the R2R3-MYB gene family in quinoa (Chenopodium quinoa) under abiotic stress
Pengcheng Ding A B C , Peng Tang A B C , Xiaofen Li A B C , Adeela Haroon D , Saima Nasreen E , Hafeez Noor A B C * , Kotb A. Attia
A B C * , Kotb A. Attia  F , Asmaa M. Abushady G H , Rongzhen Wang A B C , Kaiyuan Cui A B C , Xiangyun Wu I , Min Sun A B C * and Zhiqiang Gao A B C *
F , Asmaa M. Abushady G H , Rongzhen Wang A B C , Kaiyuan Cui A B C , Xiangyun Wu I , Min Sun A B C * and Zhiqiang Gao A B C *
A
B
C
D
E
F
G
H
I
Handling Editor: Sajid Fiaz
Abstract
The MYB transcription factor (TF) are among the largest gene families of plants being responsible for several biological processes. The R2R3-MYB gene family are integral player regulating plant primary and secondary metabolism, growth and development, and responses to hormones and stresses. The phylogenetic analysis combined with gene structure analysis and motif determination resulted in division of R2R3-MYB gene family into 27 subgroups. Evidence generated from synteny analyses indicated that CqR2R3-MYBs gene family is featured by tandem and segmental duplication events. On the basis of RNA-Seq data, the expression patterns of different tissues under salt treatment were investigated resulting CqR2R3-MYB genes high expression both in roots and stem of quinoa (Chenopodium quinoa) plants. More than half of CqR2R3-MYB genes showed expression under salt stress. Based on this result, CqR2R3-MYBs may regulate quinoa plant growth development and resistance to abiotic stresses. These findings provided comprehensive insights on role of CqR2R3-MYBs gene family members in quinoa and candidate MYB gene family members can be further studies on their role for abiotic stress tolerance in crop plants.
Keywords: CqR2R3-MYBs, function prediction, gene expression, marginal soils, nutritious crop, quinoa, stress response.
Introduction
Quinoa (Chenopodium quinoa; 2n = 4x = 36) is a nutritious crop well adapted under varying agro-ecosystems around the world. Quinoa was first domesticated approximately 7000 years ago during pre-Columbian era, and the seed was known as ‘mother grain’ during Inca Empire of 13th century (Jarvis et al. 2017). Quinoa possesses high genetic diversity and ability to be adapted under harsh conditions; i.e. marginal soils and resilient to abiotic stresses, e.g. low and high temperature, drought, salinity and large day-night duration and temperature variations (Jacobsen et al. 2005, 2007; Ruiz et al. 2014). Quinoa produces highly nutritious and gluten-free grains with an excellent balance between carbohydrates, essential amino acids, oils, minerals, vitamins and dietary fibre (Graf et al. 2015; Imamura et al. 2018). Quinoa improvement through breeding approaches has been challenging owing to multiple reasons. Therefore, genetic based interventions hold promising effect for crop improvement. Therefore, identification of candidate genes could be reliable approach can be useful technique utilised for crop improvement. The development of quinoa genome is considered useful resource for understanding functional genomics of integralagronomic attributes and the identification of candidate genes being utilised for quinoa crop improvement programmes (Jarvis et al. 2017).
Approximately all biological processes taking place in eukaryotic cells or organisms are controlled through gene expression (Pucker et al. 2020). Therefore, gene transcription regulation level is an important starting point for genetic understanding. The transcription factors (TFs) (sequence-specific DNA-bindings factor) are the proteins regulating rate of transcription of genetic information from DNA to messenger RNA through linkage with a specific DNA sequence (Sabir et al. 2022). TFs can be divided into various multigene families as per their DNA-binding domains. One of the largest TFs families present in plant kingdom has been identified as MYB gene family, present among all eukaryotes. The COLORED1 (C1) was first plant MYB gene identified in Zea mays, which was responsible for regulating anthocyanin biosynthesis both in aleurone and scutellum tissues of maize kernel (Arce-Rodríguez et al. 2021). The highly conservative MYB domain of MYB gene family members consisted of one to four neighbouring incomplete tandem repeats. Furthermore, each repeated unit is comprised of approximately 50–55 amino acid residues, with three spaced tryptophan residues, forming a helix-turn-helix (HTH) fold. The MYB TFs possess several regulatory roles, the amino acid sequence outside the conserved MYB domain are highly divergent (Zhang et al. 2020). These TFs are further divided into four classes based on the number of MYB domain repeats: 1R-MYB, R2R3-MYB, 3R-MYB and 4R-MYB (Zhao et al. 2020). The TFs have diverse functions and may play an indispensable role during plant growth and stress resistance; i.e. such as cell morphogenesis, secondary metabolism, organ morphogenesis, chloroplast development, leaf senescence, response to phosphate starvation and tolerance to cold, drought and salt stress (Park et al. 2018; Sun et al. 2018). The R2R3-MYB, TFs are considered largest subfamily among MYB family in plants whereas, the number of 4R-MYB and 3R-MYB genes are relatively smaller.
During the past decade, substantial huge amount of genome sequencing data, The R2R3-MYB gene family has been comprehensively studied for gene identification in several plant genomes; e.g. 126 genes in Arabidopsis thaliana (Stracke et al. 2001), 192 in Populus ciliata (Wilkins et al. 2009), 244 in Glycine max (Du et al. 2012) and 406 in Gossypium hirsutum (Wang et al. 2019). In addition, literature has displayed R2R3-MYB TFs role for controlling physiological and biochemical mechanisms; i.e. responses to ambient stimuli (He et al. 2020), specialised metabolism (Yan et al. 2020a), plant development (Sun et al. 2019) and cell differentiation (Yan et al. 2020b).
Previous literature has displayed involvement of R2R3-MYB gene family may play central role during plant growth and development and response of plant under salinity stress condition. However, the role of R2R3-MYB gene family toward salt stress for quinoa plant species is limited. In this study, we identified R2R3-MYB TFs their comprehensive phylogenetic analysis, chromosomal distribution, gene duplication events and synteny analysis. Based on available transcriptomic data, the tissue differential expression patterns and response of gene family under salinity stress will be explored for the characterisation of CqMYB genes and providing novel insights and valuable information for the further identification of this R2R3-MYBs gene family in quinoa.
Materials and methods
Identification and sequence analysis of the CqMYB gene family
The genomic sequences of quinoa was downloaded from the Phytozome database (http://www.phytozome.net/). The Hidden Markov Model (HMM) profile of the MYB DNA binding domain were downloaded from Pfam database (http://pfam.xfam.org/) and studied for the identification of MYB genes in the quinoa (Chenopodium quinoa Willd.) genome by utilising the simple HMM search program of TBtools (https://bio.tools/tbtools) (Chen et al. 2020). The National Center for Biotechnology Information (NCBI) (https://www.ncbi.nlm.nih.gov/) conserved domain search and SMART program (http://smart.embl-heidelberg.de/) were employed to test the presence of MYB domain. The sequence integrity of CqMYBs were analysed through multiple sequence alignment analysis by ClustalW (https://www.genome.jp/tools-bin/clustalw) (Chenna et al. 2003). The basic properties of CqMYB proteins, including protein length, isoelectric point (pI) and molecular weight (MW), were analysed by ExPasy (https://www.expasy.org/).
Phylogenetic analyses and function predictions of CqMYBs
The MYB protein sequence from quinoa and Arabidopsis thaliana L. were aligned by the ClustalW program and manually adjusted, the multiple sequence alignments were used for phylogenetic analysis. The phylogenetic tree was constructed by the neighbour-joining method employing MEGA 7.0 (https://www.megasoftware.net/) with 1000 bootstrap replicates (Kumar et al. 2016). The predictions of the biological functions of few MYB proteins were made, according to the orthology based on the aforementioned phylogenetic tree.
Chromosome distribution and gene duplication
To determine R2R3-MYBs gene family chromosomal distribution of quinoa, the annotation information were downloaded from quinoa genome from Phytozome database, the gene distribution was visualised using TBtools (Jarvis et al. 2017; Chen et al. 2020). Using multiple Collinearity Scan toolkit (https://github.com/wyp1125/MCScanX) (MCScanX) and BLASTP method (https://blast.ncbi.nlm.nih.gov/Blast.cgi?PAGE=Proteins), we also analysed the gene duplication events and their synteny relationship among different species (Chen et al. 2020). The non-synonymous (ka) and synonymous (ks) substitutions of identified gene pairs were calculated employing TBtools (Chen et al. 2020).
Gene structure and conserved motif analysis of the CqMYB gene family
The organisation of exon-intron of CqMYB gene family was illustrated and classificate through GSDS 2.07 (http://gsds.gao-lab.org/) (Hu et al. 2015). The MEME program of TBtools was utilised for identification of conserved motifs in the 103 CqMYB protein sequences. The optimised parameters of MEME were employed as follows: Number PerSeq Mode; number of motifs to find, 10; and optimum width of each motif, 6–60 residues. The MEME results were visualised by TBtools software (Chen et al. 2020).
Gene expression analysis
To characterise the expression pattern of quinoa R2R3-MYB gene family under salt stress, gene expression differences across the tissues (root, stem and leaf) were calculated from publicly available quinoa RNA-Seq datasets Sequence Read Archive (https://www.ncbi.nlm.nih.gov/sra). Materials from root (SRX8974488, SRX8974489, SRX8974490), stem (SRX8974491, SRX8974492, SRX8974493) and leaf (SRX8974494, SRX8974495, SRX8974496) were used to identify the tissue specific expression profiles. Moreover, control (SRX7700532, SRX7700537-SRX7700543) and salt stress treatment (SRX7700524-SRX7700531, SRX7700533-SRX7700536) of salt-tolerant genotype Q68 and salt-sensitive quinoa genotype Q30 were used to examine the salt stress responsive genes.
Results
Identification of R2R3MYB family genes in quinoa
We first removed redundant sequences and screened the 358 proteins containing MYB repeatsthrough HMMER analysis. Later, proteins harbouring only two MYB repeats were screened, followed by verification through Pfam and SMART. In addition, 103 quinoa R2R3-MYBs gene were identified. The identified CqR2R3-MYB genes ranged from 267 to 9033 bp, and their corresponding proteins ranged from 88 to 979 amino acids in length. The protein mass ranged from 10.08 to 110.49 kDa, whereas the isoelectric point (pI) values ranged from 4.23 to 10.26. Additionally, the subcellular localisation of CqR2R3-MYB proteins were predicted, 86.4% presented a unique nuclear localisation signal (NLS), 13.5% in addition to NLS signals for extracellular localisation were identified. Only 14 of 103 CqR2R3-MYB proteins were not present at NSL (see Supplementary Table S1).
Phylogenetic analysis of the MYB family
To explore the relationships between the R2R3-MYB proteins, we constructed the NJ-phylogenetic tree by using the amino acid sequences from both Arabidopsis and quinoa. As per classification in Arabidopsis, quinoa R2R3-MYB proteins were divided into 27 groups. Among these, group 20 was identified as largest subgroup and contained nine CqR2R3-MYB proteins (Fig. 1).
Chromosomal localisation of the CqR2R3MYB gene family
The chromosomal distribution of R2R3-MYBs gene family were analysed via employing quinoa genome annotation and TBtools was employed for their visualisation (Jarvis et al. 2017; Chen et al. 2020). The chromosomal location analysis revealed that 103 CqR2R3-MYB genes were unevenly distributed across all 18 quinoa chromosomes, whereas five CqR2R3-MYB genes belonged to unmapped scaffolds. Chromosome 1 had 14 of CqR2R3-MYB genes, followed by 12 on chromosome 6. In contrast, only a single CqR2R3-MYB genes was identified on chromosome 6 (Fig. 2).
Gene structure and protein motif analysis of the CqR2R3-MYBs
The conserved motifs of CqR2R3-MYBs were examined through employing MEME program. We identified 20 conserved motifs; however, the number of motifs (3–8) were different in identified proteins. Generally, the location of introns and exons within the genome may provide important information on evolutionary relationships of quinoa. Results also displayed that number of exons in CqR2R3-MYBs ranged from two to five; among these, mostly contained five exons (Fig. 3). This is the first time that the distribution of exons in CqR2R3-MYBs has been identified, providing valuable information on its evolutionary processes and expansion in quinoa and other species.
Gene duplication analysis of CqR2R3MYBs
Gene duplication, resulting from polyploidisation or during tandem and segmental duplication is associated with replication, is an integral component responsible for expansion of any gene family. Of the 103 genes, 89 were CqR2R3-MYBs, which accounts for 86.4% of the total genes identified that participated in gene replication events. Furthermore, 10 genes took part in tandem duplication events and 89 genes participated in WGD or segmental events, forming four tandem duplication gene pairs (Fig. 4).
Expression of CqR2R3MYBs genes in different tissues
To understand the tissue specificity and genes controlling stress response quinoa, the expression pattern of 103 CqR2R3-MYBs genes were studied. The RNA-seq data of various tissues of quinoa; i.e. stems, leaves and roots were downloaded from the SRA database (https://www.ncbi.nlm.nih.gov/sra). The FPKM values were calculated by Hisat (ver. 2.0.4, http://ccb.jhu.edu/software/hisat2/index.shtml) and Tophat software (ver. 2.1.1, http://ccb.jhu.edu/software/tophat/index.shtml). Approximately half of the CqR2R3-MYBs genes shown clear and similar tissue-specific expression (Fig. 5). CqMYB98 shown lowest expression in leaves, whereas the strongest expressions of genes were detected in the roots. In addition, the strongest expressions of CqMYB79 were found in stem. However, they showed low expression in rest of the tissues. Among the CqR2R3-MYBs genes, the most obvious differences in tissue expression were observed for genes CqMYB98, CqMYB103, CqMYB56, CqMYB95, CqMYB79 and CqMYB47.
Expression of CqMYB genes under salt treatment
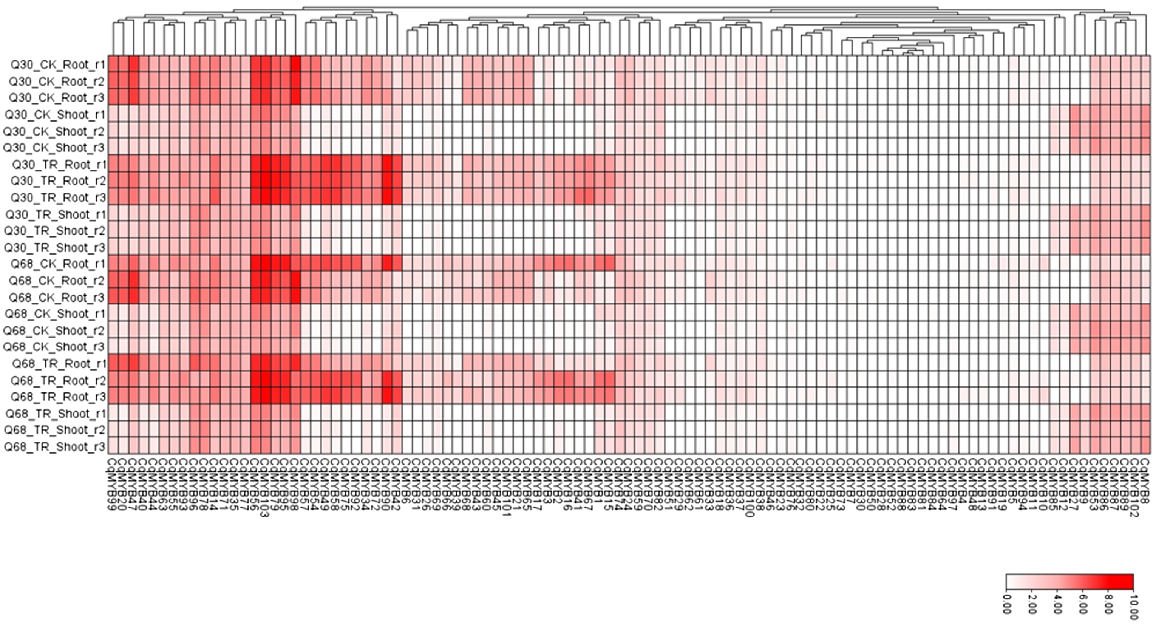
To understand the stress response of CqR2R3-MYBs, both salttolerant and salt-sensitive quinoa tissue were selected to study the expression levels of CqR2R3-MYBs under salt stress condition. The response of CqR2R3-MYBs in root were observed significantly higher compared to rest of the tissues. Approximately two thirds of genes displayed significant expression under salt stress condition, suggesting their role under salt stress condition. In roots, the expression level of CqMYB79, CqMYB95, CqMYB49, CqMYB58, CqMYB75, CqMYB92, CqMYB90, CqMYB17, CqMYB3, CqMYB2, CqMYB16, CqMYB41, CqMYB67, CqMYB1 and CqMYB15 were significantly higher for both salt-tolerant and salt-sensitive quinoa genotypes under control condition. In contrast, the expression level of CqMY-B98 was significantly lower compared to control condition for both investigated genotypes. However, in shoot, the expression levels of CqMYB78, CqMYB96, CqMYB56 and CqMYB103 were slightly higher than control condition for both salt-tolerant genotype Q68 and salt-sensitive genotype Q30 (Fig. 6).
Discussion
In this study, R2R3-MYB TPs of Arabidopsis were studied as reference genes. A total of 103 CqR2R3-MYB genes were identified in quinoa. The size of the R2R3-MYB gene family in quinoa were much higher compared to Prunus avium (69 members) (Zhao et al. 2020; Sabir et al. 2022), but lower than that Phyllostachys edulis (114 members), and Arabidopsis (125 members) (Stracke et al. 2001; Hou et al. 2018). The results published by various studies and considering phylogenetic analysis we divided the quinoa R2R3-MYB family into 27 groups. The chromosome distribution results displayed R2R3-MYB genes were unevenly distributed on 17 chromosomes. The abundance of TFs were found largely dependent on sequence duplications during genome evolution process (Yue et al. 2018). The relatively high number of CqR2R3-MYB genes in quinoa indicated that gene duplication events may have been occurred during genome evolution. It can also be hypothesised that presence of most R2R3-MYB genes in quinoa genome contain specific requirements and are involved in complicated mechanism of transcriptional regulation. These findings may provide with an insights on evolutionary relationship of R2R3-MYB family genes.
To understand the CqMYB gene family evolutionary relationships, the R2R3-MYB proteins were divided into S1-C27 groups based phylogenetic tree constructed with Arabidopsis. The CqR2R3-MYB genes grouped with Arabidopsis orthologues in several branches, suggesting R2R3-MYB function was extensively conserved across the species. The phylogenetic connection of MYB proteins between quinoa and Arabidopsis also revealed clades with varied numbers of AtMYB and CqMYB genes, which demonstrate that these two species share the similar evolutionary history. In phylogenetic tree, several CqMYB genes from quinoa were not grouped with the AtMYBs from Arabidopsis, displaying these genes were not retained between quinoa and Arabidopsis and may hold unique role. Furthermore, these results indicated the existence of divergence among primitive ancestor, species-specific MYBs genes were included in quinoa or eliminated from Arabidopsis lineages.
The genes fall in identical group contain similar positions of exon-intron, its structure; however, few dissimilarities were also observed which may be outcome of gain, loss, or skimming of introns during the evolution of MYB gene family. Similar observations were recorded for quinoa, suggesting similar evolutionary process, suggesting elimination of few genes during evolutionary process (Du et al. 2012). We showed that several genes contain only single intron, indicating the loss of intron during the evolution process. In other studies, it has been concluded that plants during evolution, gain/loss introns which is outcome of selection pressure (Wang et al. 2016; Zhao et al. 2020). The non-existence of intronic region in genes may helpful in acceleration of evolution of genes through process of gene copying (Lecharny et al. 2003). Moreover, the function of genes without an intron and those with single intron were found identical in their evolution and functional properties. We demonstrated that genes conserved through the evolutionary process contain highly functional similarities, and the majority of CqMYB genes have similar motif compositions in the identical subfamily. A sum of 20 diverse motifs were identified among MYB genes of sweet cherry (Prunus avium) with each gene containing at least two identical motifs. The arrangement and amount of the 12 distinct motif types in quinoa expressed that MYB members were functionally divergent. The presence of similar motifs and intron/exon structure among identified genes suggest their functional similarities.
A number of studies have shown that R2R3-MYB TFs play crucial roles for regulating plant growth, development and response under abiotic stresses. We investigated differential expression analysis among various quinoa tissue to salt stress, and found that five genes that were differentially expressed in any two tissues; e.g. leaf, stem and root and further divided into three groups based on difference in expression patterns. By comparing RNA-Seq data collected before and after salt stress, we identified 15 genes having significantly higher expression compared to control condition for both salt-tolerant and salt-sensitive quinoa genotypes. Furthermore, our results showed that the majority of genes were only expressed in roots, followed by four genes expressed in shoots.
Conclusions
We found 103 R2R3-MYB gene family members classified into 27 subgroups of quinoa genome. The transcriptome data showed that expression abundance of MYB family members in quinoa was significant, with CqMYB98, CqMYB103, CqMYB56, CqMYB95 and CqMYB79 genes having the high expression level and significant tissue specificity. The expression of genes under salt stress suggested members of quinoa MYB gene family are responsible for salinity stress response in quinoa. However, CqMYB90 maintained highly significant response under salt stress In conclusion, this study provided insight on functional roles of CqR2R3-MYBs, for plant growth and abiotic stress. These results will be useful for our enhancing understanding on molecular mechanisms of stress tolerance in crops, especially quinoa.
Data availability
The data used to support the findings of this research are available from the corresponding author upon request.
Declaration of funding
This paper was financed by The Global Crop Diversity This research was supported by the Research Program Sponsored by Ministerial and Provincial Co-Innovation Centre for Endemic Crops Production with High-quality and Efficiency in Loess Plateau, Taigu 030801, China (No. SBGJXTZX-15), the National Modern Agricultural Industrial Technology System (CARS-03-01-24), the Key Laboratory of Crop Ecology and Dryland Cultivation Physiology of Shanxi Province (201705D111007), the Key Innovation Team of the 1331 Project of Shanxi Province, the Scientific and Technological Innovation Project of colleges in Shanxi Province (2021L178), Science and technology innovation fund of Shanxi Agricultural University (2018YJ18).
Author contributions
Zhiqiang Gao conceived the study design; Pengcheng Ding collected data; Peng Tang, Xiaofen Li, Hafeez Noor, Kaiyuan Cui, Rongzhen Wang and Xiangyun Wu performed the statistical analyses, interpreted the results and drafted the manuscript. Hafeez Noor, Min Sun, Pengcheng Ding, Kotb A. Attia, Asmaa M. Abushady, Adeela Haroon, Saima Nasreen, Rongzhen Wang and Peng Tang performed a critical revision of the manuscript. All authors approved the version to be published and agreed to be accountable for all aspects of the work.
Acknowledgements
The authors acknowledge Researchers Supporting Project number (RSP-2024R369), King Saud University, Riyadh, Saudi Arabia. This research was supported by the Research Program Sponsored by Ministerial and Provincial Co-Innovation Centre for Endemic Crops Production with High-quality and Efficiency in Loess Plateau, Taigu 030801, China (No. SBGJXTZX-15), the National Modern Agricultural Industrial Technology System (CARS-03-01-24), the Key Laboratory of Crop Ecology and Dryland Cultivation Physiology of Shanxi Province (201705D111007), the Key Innovation Team of the 1331 Project of Shanxi Province, the Scientific and Technological Innovation Project of colleges in Shanxi Province (2021L178), Science and technology innovation fund of Shanxi Agricultural University (2018YJ18).
References
Arce-Rodríguez ML, Martínez O, Ochoa-Alejo N (2021) Genome-wide identification and analysis of the MYB transcription factor gene family in chili pepper (Capsicum spp.). International Journal of Molecular Sciences 22(5), 2229.
| Crossref | Google Scholar | PubMed |
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, et al. (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Molecular Plant 13, 1194-1202.
| Crossref | Google Scholar | PubMed |
Chenna R, Sugawara H, Koike T, Lopez R, Gibson TJ, Higgins DG, et al. (2003) Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Research 31, 3497-3500.
| Crossref | Google Scholar | PubMed |
Du H, Yang S-S, Liang Z, et al. (2012) Genome-wide analysis of the MYB transcription factor superfamily in soybean. BMC Plant Biology 12, 106.
| Crossref | Google Scholar | PubMed |
Graf BL, Rojas-Silva P, Rojo LE, Delatorre-Herrera J, Baldeón ME, Raskin I (2015) Innovations in health value and functional food development of quinoa (Chenopodium quinoa Willd.). Comprehensive Reviews in Food Science and Food Safety 14, 431-445 10.1111/1541-4337.12135.
| Google Scholar |
He J, Liu Y, Yuan D, et al. (2020) An R2R3 MYB transcription factor confers brown planthopper resistance by regulating the phenylalanine ammonia-lyase pathway in rice. Proceedings of the National Academy of Sciences 117, 271-277.
| Crossref | Google Scholar |
Hou D, Cheng Z, Xie L, Li X, Li J, Mu S, Gao J (2018) The R2R3MYB gene family in Phyllostachys edulis: genome-wide analysis and identification of stress or development-related R2R3MYBs. Frontiers in Plant Science 9, 738.
| Crossref | Google Scholar | PubMed |
Hu B, Jin J, Guo A-Y, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31, 1296-1297.
| Crossref | Google Scholar | PubMed |
Imamura T, Takagi H, Miyazato A, Ohki S, Mizukoshi H, Mori M (2018) Isolation and characterization of the betalain biosynthesis gene involved in hypocotyl pigmentation of the allotetraploid Chenopodium quinoa. Biochemical and Biophysical Research Communications 496(2), 280-286.
| Crossref | Google Scholar | PubMed |
Jacobsen S-E, Monteros C, Christiansen JL, Bravo LA, Corcuera LJ, Mujica A (2005) Plant responses of quinoa (Chenopodium quinoa Willd.) to frost at various phenological stages. European Journal of Agronomy 22, 131-139.
| Crossref | Google Scholar |
Jacobsen S-E, Monteros C, Corcuera LJ, Bravo LA, Christiansen JL, Mujica A (2007) Frost resistance mechanisms in quinoa (Chenopodium quinoa Willd.). European Journal of Agronomy 26, 471-475.
| Crossref | Google Scholar |
Jarvis DE, Ho YS, Lightfoot DJ, Schmöckel SM, Li B, Borm TJA, Ohyanagi H, Mineta K, Michell CT, Saber N, et al. (2017) The genome of Chenopodium quinoa. Nature 542, 307-312.
| Crossref | Google Scholar | PubMed |
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution 33, 1870-1874.
| Crossref | Google Scholar | PubMed |
Lecharny A, Boudet N, Gy I, Aubourg S, Kreis M (2003) Introns in, introns out in plant gene families: a genomic approach of the dynamics of gene structure. Journal of Structural and Functional Genomics 3(1–4), 111-116.
| Crossref | Google Scholar | PubMed |
Park D-Y, Shim Y, Gi E, Lee B-D, An G, Kang K, et al. (2018) The MYB-related transcription factor RADIALIS-LIKE3 (OsRL3) functions in ABA-induced leaf senescence and salt sensitivity in rice. Environmental and Experimental Botany 156, 86-95.
| Crossref | Google Scholar |
Pucker B, Pandey A, Weisshaar B, Stracke R (2020) The R2R3-MYB gene family in banana (Musa acuminata): genome-wide identification, classification and expression patterns. PLoS ONE 15(10), e0239275.
| Crossref | Google Scholar | PubMed |
Ruiz KB, Biondi S, Oses R, et al. (2014) Quinoa biodiversity and sustainability for food security under climate change. A review. Agronomy for Sustainable Development 34, 349-359.
| Crossref | Google Scholar |
Sabir IA, Manzoor MA, Shah IH, Liu X, Zahid MS, Jiu S, Wang J, Abdullah M, Zhang C (2022) MYB transcription factor family in sweet cherry (Prunus avium L.): genome-wide investigation, evolution, structure, characterization and expression patterns. BMC Plant Biology 22(1), 2.
| Crossref | Google Scholar | PubMed |
Stracke R, Werber M, Weisshaar B (2001) The R2R3-MYB gene family in Arabidopsis thaliana. Current Opinion in Plant Biology 4(5), 447-456.
| Crossref | Google Scholar | PubMed |
Sun X, Matus JT, Wong DCJ, Wang Z, Chai F, Zhang L, et al. (2018) The GARP/MYB-related grape transcription factor AQUILO improves cold tolerance and promotes the accumulation of raffinose family oligosaccharides. Journal of Experimental Botany 69, 1749-1764.
| Crossref | Google Scholar | PubMed |
Sun W, Gao Z, Wang J, et al. (2019) Cotton fiber elongation requires the transcription factor GhMYB212 to regulate sucrose transportation into expanding fibers. New Phytologist 222, 864-881.
| Crossref | Google Scholar | PubMed |
Wang M, Yue H, Feng K, Deng P, Song W, Nie X (2016) Genome-wide identification, phylogeny and expressional profiles of mitogen activated protein kinase kinase kinase (MAPKKK) gene family in bread wheat (Triticum aestivum L.). BMC Genomics 17(1), 668.
| Crossref | Google Scholar |
Wang N, Ma Q, Ma J, et al. (2019) A comparative genome-wide analysis of the R2R3-MYB gene family among four gossypium species and their sequence variation and association with fiber quality traits in an interspecific G. hirsutum × G. barbadense population. Frontiers in Genetics 10, 741.
| Crossref | Google Scholar |
Wilkins O, Nahal H, Foong J, Provart NJ, Campbell MM (2009) Expansion and diversification of the Populus R2R3-MYB family of transcription factors. Plant Physiology 149, 981-993.
| Crossref | Google Scholar | PubMed |
Yan S, Chen N, Huang Z, et al. (2020a) Anthocyanin fruit encodes an R2R3-MYB transcription factor, SlAN2-like, activating the transcription of SlMYBATV to fine-tune anthocyanin content in tomato fruit. New Phytologist 225, 2048-2063.
| Crossref | Google Scholar | PubMed |
Yan Y, Li C, Dong X, et al. (2020b) MYB30 is a key negative regulator of Arabidopsis photomorphogenic development that promotes PIF4 and PIF5 protein accumulation in the light. The Plant Cell 32, 2196-2215.
| Crossref | Google Scholar | PubMed |
Yue H, Shu D, Wang M, Xing G, Zhan H, Du X, Song W, Nie X (2018) Genome-wide identification and expression analysis of the HD-zip gene family in wheat (Triticum aestivum L.). Genes 9(2), 70.
| Crossref | Google Scholar | PubMed |
Zhang Y, Xu S, Cheng Y, Wang J, Wang X, Liu R, et al. (2020) Functional identification of PsMYB57 involved in anthocyanin regulation of tree peony. BMC Genetics 21, 124.
| Crossref | Google Scholar | PubMed |
Zhao K, Cheng Z, Guo Q, Yao W, Liu H, Zhou B, Jiang T (2020) Characterization of the poplar R2R3-MYB gene family and over-expression of PsnMYB108 confers salt tolerance in transgenic tobacco. Frontiers in Plant Science 11, 571881.
| Crossref | Google Scholar | PubMed |