Identification and expression analysis of SBP-Box-like (SPL) gene family disclose their contribution to abiotic stress and flower budding in pigeon pea (Cajanus cajan)
Tayyaba Shaheen A , Abdul Rehman A , Amany H. A. Abeed B , Muhammad Waqas A , Asad Aslam C , Farrukh Azeem
A , Amany H. A. Abeed B , Muhammad Waqas A , Asad Aslam C , Farrukh Azeem  A * , Muhammad Qasim A , Muhammad Afzal A , Muhammad Farooq Azhar D , Kotb A. Attia
A * , Muhammad Qasim A , Muhammad Afzal A , Muhammad Farooq Azhar D , Kotb A. Attia  E , Asmaa M. Abushady F G , Sezai Ercisli H and Nazia Nahid
E , Asmaa M. Abushady F G , Sezai Ercisli H and Nazia Nahid  A *
A *
A
B
C
D
E
F
G
H
Abstract
The SPL gene family (for Squamosa Promoter-binding like Proteins) represents specific transcription factors that have significant roles in abiotic stress tolerance, development and the growth processes of different plants, including initiation of the leaf, branching and development of shoot and fruits. The SPL gene family has been studied in different plant species; however, its role is not yet fully explored in pigeon pea (Cajanus cajan). In the present study, 11 members of the CcSPL gene family were identified in C. cajan. The identified SPLs were classified into nine groups based on a phylogenetic analysis involving SPL protein sequences from C. cajan, Arabidopsis thaliana, Cicer arietinum, Glycine max, Phaseolus vulgaris, Vigna unguiculata and Arachis hypogaea. Further, the identification of gene structure, motif analysis, domain analysis and presence of cis-regulatory elements in the SPL family members were studied. Based on RNA-sequencing data, gene expression analysis was performed, revealing that CcSPL2.1, 3 and 13A were significantly upregulated for salt-tolerance and CcSPL14 and 15 were upregulated in a salt-susceptible cultivar. Real-time qPCR validation indicated that CcSPL3, 4, 6 and 13A were upregulated under salt stress conditions. Therefore, molecular docking was performed against the proteins of two highly expressed genes (CcSPL3 and CcSPL14) with three ligands: abscisic acid, gibberellic acid and indole-3-acetic acid. Afterward, their binding affinity was obtained and three-dimensional structures were predicted. In the future, our study may open avenues for harnessing CcSPL genes in pigeon pea for enhanced abiotic stress resistance and developmental traits.
Keywords: bioinformatics, genome-wide analysis, molecular docking, molecular interaction analysis, next-generation sequencing, omics data analysis, plant hormone, salinity stress, structure prediction.
Introduction
The pigeon pea (Cajanus cajan (L.) Millsp.) is a diploid (2n = 22) legume from the tribe Phaseoleae positioned in the millettioid (warm-season legume) clade within the sub-family Papilionoidea. It is grown in humid and subtropic regions around the world (Asia, Africa and Central/South America). Pigeon pea is cultivated on more than 5 million ha, making it the sixth-most vital leguminous plant in the world. It is historically important and is mainly cultivated in Asian subcontinent countries like Pakistan, India and Bangladesh. Approximately 85% of the world’s pigeon pea is produced in India. It is mainly used as a food resource and has high nutritional value, representing a privileged protein source for more than one billion people around the globe. Pigeon pea is cultivated in low-input and risk-prone bordering surroundings (Dar 2009). Genome sequencing revealed that pigeon pea potentially has ~48 000 genes, which are an important genetic source for various traits of plants (Varshney et al. 2012; Noman and Azhar 2023). For instance, the drought tolerance-related gene families have played a vital role throughout the evolution of the pigeon pea (Varshney et al. 2012). In the NCBI database (National Center for Biotechnology Information, https://www.ncbi.nlm.nih.gov/), pigeon pea contains one reference genome (C. cajan_V1.0) containing a genome size of approximately 592.8 Mb. The contig N50 is approximately 22.5 kb and contains 33 495 genes. In the current genome assembly, 29 220 (87.2) are protein-coding genes, 1838 (5.5%) non-coding genes, 1247 (3.7%) small RNAs and 1187 (3.5%) pseudogenes (Varshney et al. 2012). Despite having excellent stress tolerance features, very few studies have been devised to explore potential stress tolerance mechanisms in this crop.
Squamosa Promoter-binding like Proteins (SBP-Box or SPL) are among the main transcription factors that play an important role in the development and determination of plant architecture (Preston and Hileman 2013). The SPL proteins contain a common DNA-binding domain (SBP) with a remarkable zinc-finger domain pattern. SPL genes are also involved in gibberellic acid (GA3) signalling, which controls some of the important aspects of plant growth like germination of the seeds, flowering and the elongation/growth of the plants, as well as the toxin response in plants (Ahmed et al. 2021). Three main motifs were predicted in the SPL gene family domain (Yang et al. 2008). The motif includes two Zinc-binding sites (Zn-1 and Zn-2). The first Zinc-binding site (Zn-1) was Cysteine-Cysteine-Cysteine-Histidine and the second Zinc-binding site (Zn-2) was Cysteine-Cysteine-Histidine-Cysteine. A Nuclear Localisation Signal (NLS) was also present, as a Lysine-Arginine complex (Palmeri and Malim 1999). The SPL gene family has an important role in the major biological and physiological processes of growth and development of plants (Hou et al. 2013; Waqas et al. 2019; Nadeem et al. 2023).
The SPL transcription factors control several features of plant development and physiology. The 17 SPL members of Arabidopsis thaliana genome are classified into Class I members, comprising large proteins of more than 800 amino acids (SPL1, 7, 12, 14 and 16) and Class II (remaining 12 SPLs) members, comprising smaller proteins (up to 400 amino acids) (Cardon et al. 1999). The SPL-II members are targets of miR156 (except SPL8) and are involved in plant phase transitions (Wu and Poethig 2006; Wu et al. 2009; Yu et al. 2010; Xue et al. 2014). SPL8 plays a key role in pollen sac development. SPL2 and SPL11 are downregulated in response to heat stress and are involved in heat-shock memory (Stief et al. 2014). In Chlamydomonas reinhardtii (a green alga), the Class I SPL homologue CRR1 regulates signalling related to copper nutrition and performs transcriptional activation of genes in response to copper deficiency (Kropat et al. 2005; Abeed et al. 2021). Excitingly, SPL7 (Class I) also plays a key role in copper homeostasis and response to light. SPL14 is important for plant architecture and biotic stress response (to the fungal toxin fumonisin B1) (Jorgensen and Preston 2014). The involvement of SPLs in a diverse range of plant processes motivates us to explore their role in more detail.
The SPLs have been reported in several plant species of economic importance (Cao et al. 2019; Zhong et al. 2019; Yu et al. 2021; Sun et al. 2022). However, to our knowledge, no study has reported on the SPL gene family in C. cajan. The primary objectives of this study were to explore and characterise the members of the SPL gene family in pigeon pea (C. cajan) and to gain insights into their functions. Specifically, we aimed to identify and classify the CcSPL genes, analyse their gene structures, motifs, and the presence of cis-regulatory elements. Additionally, we sought to investigate the gene expression patterns of CcSPL genes in response to salt stress using RNA sequencing (RNA-Seq) data, followed by quantitative real-time polymerase chain reaction (qPCR) validation. Moreover, we conducted molecular docking studies to assess the binding affinity of two highly expressed genes, CcSPL3 and CcSPL14, with three important phytohormones, i.e. abscisic acid (ABA), gibberellic acid (GA3) and indole-3-acetic acid (IAA). The ultimate objective was to provide a foundation for a deeper understanding of the functional roles of the SPL gene family in C. cajan, especially in the context of abiotic stress tolerance and growth processes (Goddard et al. 2005).
Materials and Methods
Plant material, treatments and growth conditions
For 2 weeks, Cajanus cajan (L.) Millsp. seedlings were grown in plastic pots (3–4 seedlings/pot) filled with peat moss, in a growth chamber at a humidity level of 60% and a temperature of 22°C at night and 25°C in the day. The plants were exposed to light for 16 h, followed by 8 h of darkness. Hoagland solution (10 mL/day) was applied to provide the necessary nutrients for plant growth. The seedlings were given 100 mM NaCl for the treatment of salt stress after 2 weeks of germination. The Hoagland solution was used as a preliminary treatment for the plants. The drought-stressed plants were deprived of water supply for 7 days whereas control plants were given Hoagland solution. For RNA extraction, the plants were divided into three replicates and each replicate contained three plants. The samples were frozen immediately using liquid nitrogen and then kept at −80°C until further experimentation.
Genome-wide identification and phylogenetic analysis of SPL family in C. cajan
A total of 11 members of the SPL gene family in A. thaliana (AtSPLs) were identified from the TAIR database (https://www.arabidopsis.org/). The AtSPL protein sequences were used as a query to identify the homologues in C. cajan (CcSPLs) using BLASTp (Protein BLAST: search protein databases using a protein query, nih.gov) (Swarbreck et al. 2007). The retrieved sequences were checked for redundancy using the SIAS tool by the Immunomedicine group (Immunomedicine Group: Tools >> SIAS, ucm.es) and annotated for further analysis using atom (https://atom.io/). The results were manually verified using NCBI and Uniprot (https://www.uniprot.org/) (Sherry et al. 2001; The UniProt Consortium 2015). After investigating the protein sequences retrieved from different species, phylogenetic analysis was performed (Pasquier et al. 2016). The maximum likelihood tree was constructed using IQ Tree (http://iqtree.cibiv.univie.ac.at).
Characteristics of SPL protein
The physiochemical characteristics of SPL gene family proteins were predicted using an online tool developed by the Swiss Institute of Bioinformatics called ProtParam (https://web.expasy.org/protparam/). Motifs of the SPL gene family were identified using MEME (Multiple Em for Motif Elicitation) (https://meme-suite.org/meme/tools/meme) where site distribution was zero or one sequence per occurrence and number of motifs was selected to 10. Motif logos were also designed by MEME (https://meme-suite.org/meme/tools/meme) (Bailey et al. 2015). The subcellular localisation of proteins was projected using an online-based soft berry tool called ProtComp 9.0 (ProtComp, softberry.com). SPL protein domains were predicated using InterPro (https://www.ebi.ac.uk/interpro) and were aligned using DNAMAN software (https://www.lynnon.com/; Wang 2015).
Gene sequence analysis of CcSPL
Exon and intron structures of CcSPL and AtSPL genes were predicated using GSDS Gene Structure Display Server Tool (ver. 2.0, http://gsds.gao-lab.org/) (Hu et al. 2015).
Analysis of putative promoter region of CcSPL genes
The 2000 bp promoter and UTR (untranslated region) were identified using the NCBI genomics tool (https://www.ncbi.nlm.nih.gov/nuccore) and the cis-elements in CcSPL gene promoter regions were identified using an online server-based website called PLANT CARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/; Lescot et al. 2002).
Expression profile from RNA-seq data
To study the protein expression, the transcriptomic data on abiotic stress (SRP104063) and flower budding (SRP262441) were retrieved from the NCBI SRA (Sequence Read Archive) database (https://www.ncbi.nlm.nih.gov/sra). All the available runs were mapped to C. cajan using Galaxy Bowtie2 (map reads against reference genome) (https://usegalaxy.org/). A tool of Galaxy called Cufflinks was used to calculate gene expression levels based on FPKM values (fragments per kilobase of transcript per million mapped reads, https://usegalaxy.org/). The expression profile of both abiotic stress and flower budding experiment was incorporated using the software TBTOOLS (https://bio.tools/tbtools; Chen et al. 2020).
RNA extraction, cDNA synthesis and qPCR
Thermo Nanodrop 2000 (Thermo Fisher Scientific, Waltham, MA, USA) was used to measure the amount of total RNA, extracted from C. cajan leaf samples using the Trizol technique. The Maxima H-minus First-Strand Synthesis kit was used to synthesise complementary DNA (cDNA) from 1 μg of RNA. To prepare it for future analysis, cDNA was kept at −20°C. A qPCR analysis was carried out using an iTaq Universal SYBR Green Super-Mix and a qPCR detection system (CFX96 TouchTM RT PCR Detection System, Bio_Rad Labs, Hercules, CA, USA). Gene-specific primers were designed using the internet tool ‘Oligo Calculator’ (http://mcb.berkeley.edu/labs/krantz/tools/oligocalc.html/, accessed on 13 July 2022), which were then confirmed by the NCBI-primer BLAST algorithm (https://www.ncbi.nlm.nih.gov/tools/primer-blast/, accessed on 13 July 2022). Each gene’s expression was analysed in triplicate, and the CcGAPDH gene served as the housekeeping gene (Ali et al. 2018).
CcSPL protein interaction analysis
To study the protein–protein interaction a web-based tool called string (https://string-db.org/) was used. All the sequences of CcSPL protein were pasted into the search box one by one and the results compiled manually using Microsoft PowerPoint into a single file (Bultan et al. 2017).
3D protein structure prediction and docking analysis
Protein docking analysis of the CcSPL proteins was undertaken using multiple tools. The first was to predict the three dimensional (3D) structures of the proteins. For most of the proteins of the family, 3D structures were predicted using an online server-based tool I-TASSER (Iterative Threading ASSEmbly Refinement) (https://zhanggroup.org/I-TASSER/). CcSPL3 was predicted using QUARK, because it contains less than 200 amino acids (https://zhanggroup.org/QUARK/). QUARK proposes developing an algorithm for ab initio protein structure prediction and protein–peptide folding, to construct the right protein 3D model using only the amino acid sequence (Waqas et al. 2021). To study the protein–ligand interaction, the three ligands, i.e. ABA, GA3 and IAA, were used and were retrieved from PubChem (https://pubchem.ncbi.nlm.nih.gov/). To perform molecular docking on the predicted SPL proteins, PyRx was employed. The best docking results (with the lowest values) were selected based on the binding affinity of the molecules, and the results were visualised using software called Chimera (Kim et al. 2019; Halim et al. 2021).
Ethical approval
This article does not contain any studies with humans or animals performed by any of the authors. We sought appropriate permission from the farm or field owner during specimens’ collection and experimentation. We confirm that during the collection and execution of the experiment, the authors have complied with the International Union for Conservation of Nature (IUCN) Statement on Research Involving Species at Risk of Extinction and the Convention on the Trade in Endangered Species of Wild Fauna and Flora. All methods were performed in accordance with the relevant guidelines/regulations/legislation.
Results
Identification and phylogenetic analysis of CcSPL genes
A total of 11 SPL genes were identified in C. cajan (Table 1). The identified genes were manually curated for accession numbers, gene IDs, physiochemical properties of protein sequences, domains and subcellular localisation. The amino acid sequence length of CcSPLs ranged from 125 (CcSPL4) to 1000 (CcSPL15). All the proteins were located in the nucleus and the theoretical isoelectric points (pI) ranged from 6.04 (CcSPL1) to 9.45 (CcSPL15). The Grand Average of Hydropathicity (GRAVY) for each SPL protein was less than 0 (Liu et al. 2019). A maximum likelihood tree was constructed using IQ Tree (http://iqtree.cibiv.univie.ac.at) based on 79 SPL gene family sequences from C. cajan, A. thaliana, Cicer arietinum, Glycine max, Phaseolus vulgaris and Vigna unguiculata. All members of the tree were placed into nine groups (G1–G9: Fig. 1). Each group has at least one of the 11 members of the CcSPL protein (Nguyen et al. 2015). Two paralogous gene pairs were identified in G. max, including CcSPL1/GmSPL1 (G1) and CcSPL8/GmSPL8 (G5). For P. vulgaris, one paralogous pair CcSPL3/PvSPL3.1 was identified in G4. Similarly, a paralogous pair CcSPL13A/VuSPL13A was identified in G9 and CcSPL6/AtSPL6 in G8 for V. unguiculata and A. thaliana, respectively. In G9, CcSPL2.1/CcSPL2.2 was also identified as a paralogous pair from C. cajan. Gene pair CcSPL15/AhSPL15 in G6 was an out-paralog in the context of VuSPL15.1/15.2, GmSPL15, PvSPL15, and in paralogs for CaSPL15.
| Sr. No | Gene ID | Gene name | Protein ID | No. of amino acids | Molecular weight | Theoretical pI | Domains | GRAVY | Subcellular localisation | |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | LOC109809856 | CcSPL1 | XP_020228850.1 | 990 | 110 216.24 | 6.04 | SBP_dom, SBP, ZF_SBP | −0.428 | Nuclear | |
| 2 | LOC109809676 | CcSPL2.2 | XP_029129944.1 | 448 | 49 114.55 | 7.53 | SBP_dom, SBP, ZF_SBP | −0.694 | Nuclear | |
| 3 | LOC109814571 | CcSPL3 | XP_020234618.1 | 138 | 16 082.56 | 6.39 | SBP_dom, SBP, ZF_SBP | −1.480 | Nuclear | |
| 4 | LOC109810441 | CcSPL4 | XP_020229499.1 | 125 | 14 724.52 | 9.40 | SBP_dom, SBP, ZF_SBP | −1.256 | Nuclear | |
| 5 | LOC109808214 | CcSPL6 | XP_020226701.1 | 440 | 48 374.81 | 7.58 | SBP_dom, SBP, ZF_SBP | −0.617 | Nuclear | |
| 6 | LOC109815860 | CcSPL7 | XP_020236266.1 | 756 | 84 302.04 | 6.36 | SBP_dom, SBP, ZF_SBP | −0.293 | Nuclear | |
| 7 | LOC109788859 | CcSPL8 | XP_020203285.1 | 332 | 36 601.35 | 8.86 | SBP_dom, SBP, ZF_SBP | −0.797 | Nuclear | |
| 8 | LOC109810682 | CcSPL2.1 | XP_020229799.1 | 448 | 49 907.02 | 6.32 | SBP_dom, SBP, ZF_SBP | −0.751 | Nuclear | |
| 9 | LOC109797843 | CcSPL13A | XP_020213579.1 | 373 | 41 033.83 | 8.88 | SBP_dom, SBP, ZF_SBP | −0.650 | Nuclear | |
| 10 | LOC109816797 | CcSPL14 | XP_020237510.1 | 1000 | 111 145.24 | 8.29 | SBP_dom, SBP, ZF_SBP | −0.531 | Nuclear | |
| 11 | LOC109793190 | CcSPL15 | XP_020208244.1 | 367 | 39 169.60 | 9.45 | SBP_dom, SBP, ZF_SBP | −0.469 | Nuclear |
pI, isoelectric point; GRAVY, grand average of hydropathicity.
Unrooted phylogenetic tree of SPL protein from Cajanus cajan (Cc), Arabidopsis thaliana (At), Cicer arietinum (Ca), Glycine max (Gm), Phaseolus vulgaris (Pv), Vigna unguiculata (Vu) and Arachis hypogaea (Ah). The tree was generated using IQ Tree with the maximum likelihood method. Numbers above the branches show bootstrap value. Different colour circles represent different plants.
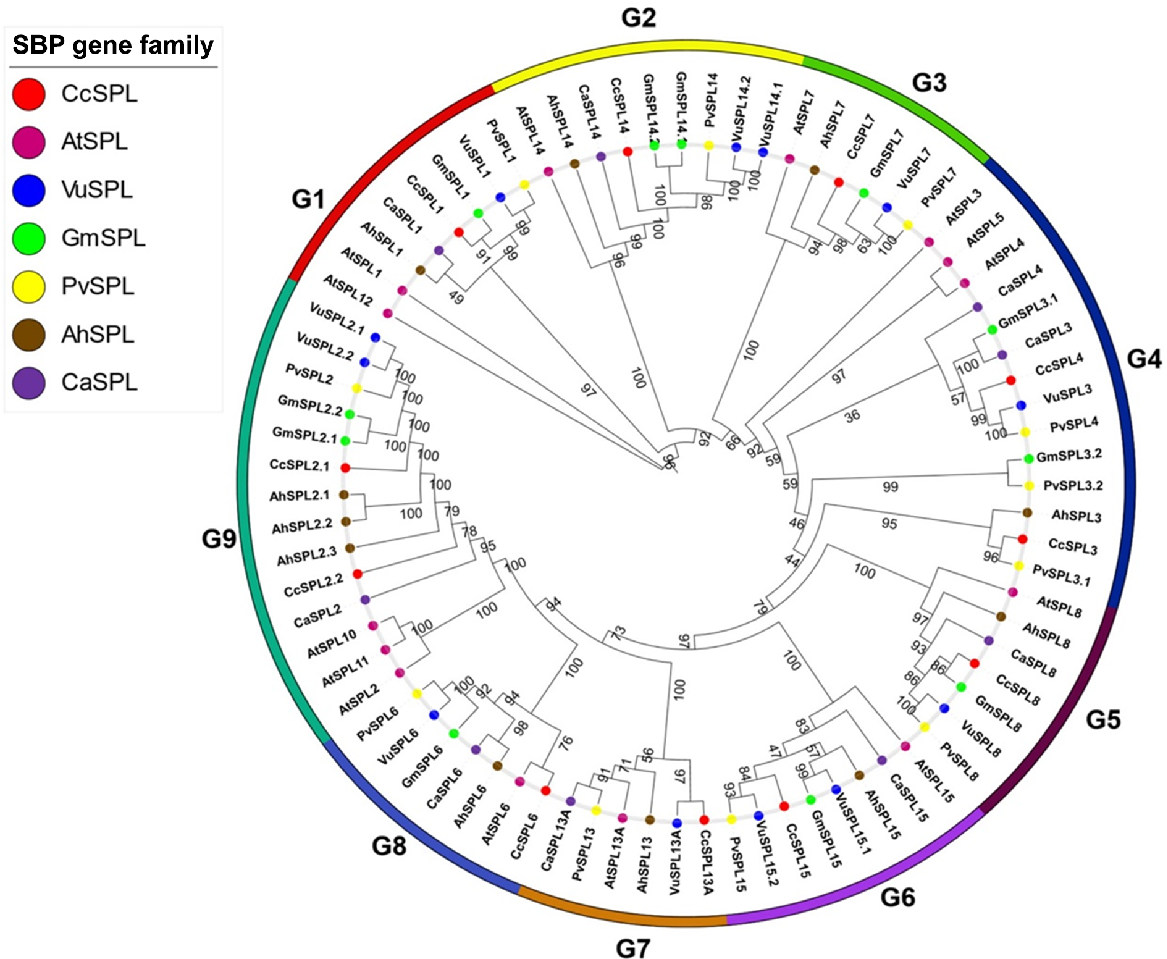
In G2, CcSPL14 was identified as orthologous to members including GmSPL14.1/14.2, PvSPL14, VuSPL14.1/14.2; and CcSPL7 in G3 was orthologous to GmSPL7/PvSPL7/VuSPL7. In G4, CcSPL4 was identified as orthologous to VuSPL3 and PvSPL4 (Yang et al. 2008).
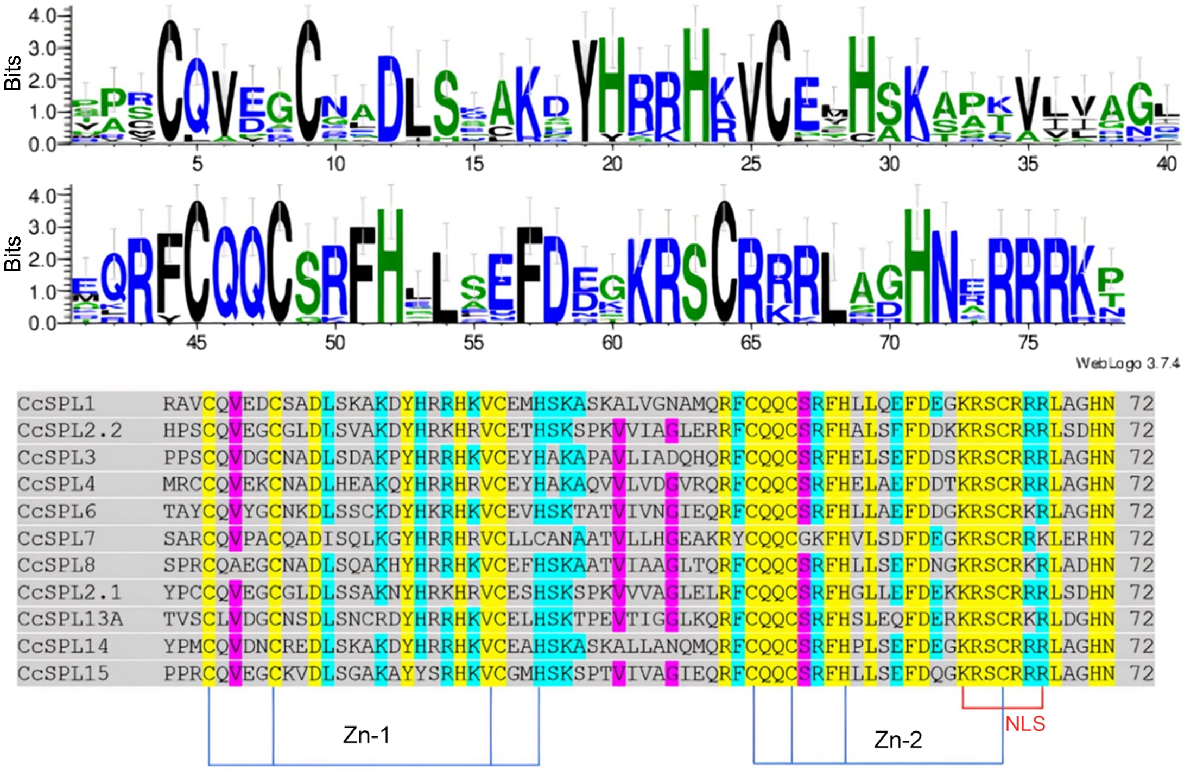
A sequence alignment of these motif regions was performed for SPL proteins and represented as a sequence logo (Fig. 2). The Zn-1 (Cys-Cys-Cys-His) was conserved in all CcSPLs except CcSPL7, where the ‘His’ was replaced with another ‘Cys’. The second Zn-binding site (Zn-2) was conserved (Cys-Cys-His-Cys) among all the CcSPL proteins. Moreover, the Nuclear Localisation Signal (NLS) was highly conserved among all the c protein sequences (Ioannidi et al. 2016).
Sequence alignment to identify conserved motifs. Multiple sequence alignment of CcSPL proteins was done using t-coffee and the logo was designed using Weblogo (Crooks et al. 2004).
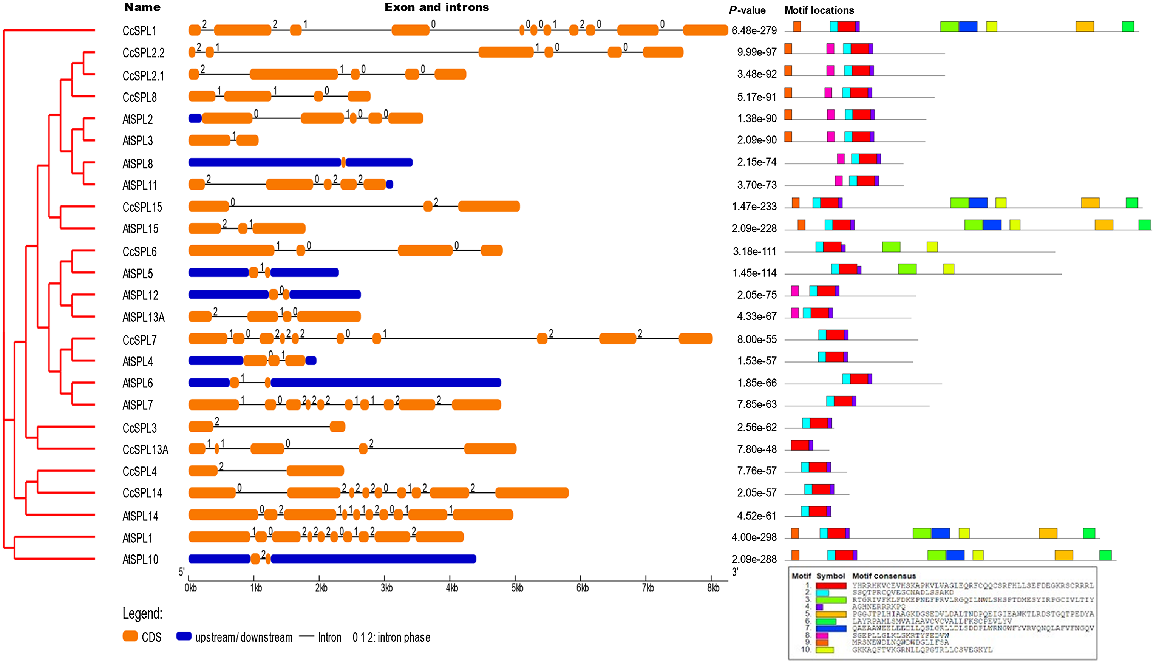
Gene structure analysis for exon/intron arrangement
Gene structures for exons and introns are shown in Fig. 3. The introns and exons of CcSPL are grouped according to phylogenetic analysis. The number of introns were not the same in all genes of the family as they ranged from 1 to 10. For instance, CcSPL1 contains 10, CcSPL7 contains 9, CcSPL14 contains 8, CcSPL2.2 contains 5, CcSPL2.1 and CcSPL13A contain 4, CcSPL6 and CcSPL8.1 contain 3, CcSPL15 contains 2, and CcSPL4 and CcSPL5 contain only one intron, respectively. Furthermore, 17 members of AtSPL and 11 members of CcSPL were analysed for the presence of conserved motifs using MEME software (ver. 5.4.1). Ten conserved motifs were detected and shown in Fig. 3. Motif-1 and Motif-4 were highly conserved in all of the proteins of the family. Motif-2 was also conserved in almost all of the genes except CcSL4. Motif-9 was also highly conserved and present in AtSPL1, AtSPL12, CcSPL1, CcSPL14, AtSPL14, AtSPL7, CcSPL7, CcSPL2.2, CcSPL2.1, AtSPL2, AtSPL10 and AtSPL11. Motif-3, 5, 6, 7 and 10 were conserved in AtSPL1, AtSPL12, CcSPL1, CcSPL14, AtSPL14 and AtSPL7. Motif-8 was conserved in CcSPL2.2, CcSPL2.1, AtSPL2, AtSPL10, AtSPL11, CcSPL15, CcSPL8, AtSPL8 and AtSPL15 (Fig. 3).
Exon–intron structures were characterised using GSDS 2.0, as shown in Fig. 3. Exons or CDS (coding sequence) are represented by yellow rectangles with rounded corners, introns by black lines, and untranslated portions by blue rectangles (UTRs). Intron stages are represented by 0, 1 and 2. Motifs are also displayed in this figure.
Protein–protein interactions analysis of CcSPL
Protein–protein interactions analysis was studied using a web-based tool String (Fig. 4a). The associations of SPL proteins with other proteins predicted through this tool are meant to be specifically significant (Takahashi et al. 2000). No interaction was observed for CcSPL2.1 and CcSPL2.2. CcSPL1 interacts with PEX4 (involved in peroxisome biogenesis) and AT2G34990 (involved in zinc ion binding). CcSPL14 interacts with PEX4 (involved in peroxisome biogenesis) and AGL20 (involved in transcription factor activity), which interacts with CcSPL3, CcSPL4, CcSPL8, CcSPL13A and CcSPL15.In contrast, AGL8 and CcSPL7 interact with COPT1, COPT2 and COPT6 (proteins involved in copper ion transmembrane transporter activity), FHY3 and HY5. Interestingly, CcSPL3, CcSPL4, CcSPL6, CcSPL8, CcSPL13A and CcSPL15 are working in a closer relationship whereas, all were interacting with the protein AGL8 (a probable transcription factor that helps in early floral meristem identity) in a complex network, which can be seen in the interaction diagram (Fig. 4).
(a) Using the online server-based technology called STRING, the protein–protein interactions of the SPL proteins with other members of the same family and other proteins. (b) Regulatory cis-elements were predicted for eight members of CcSPL family for 2000 bp promoter and UTR regions. The distribution of cis-elements related to various stress and hormone responses is depicted. Distinct shapes and colours are used to symbolise different cis-elements.
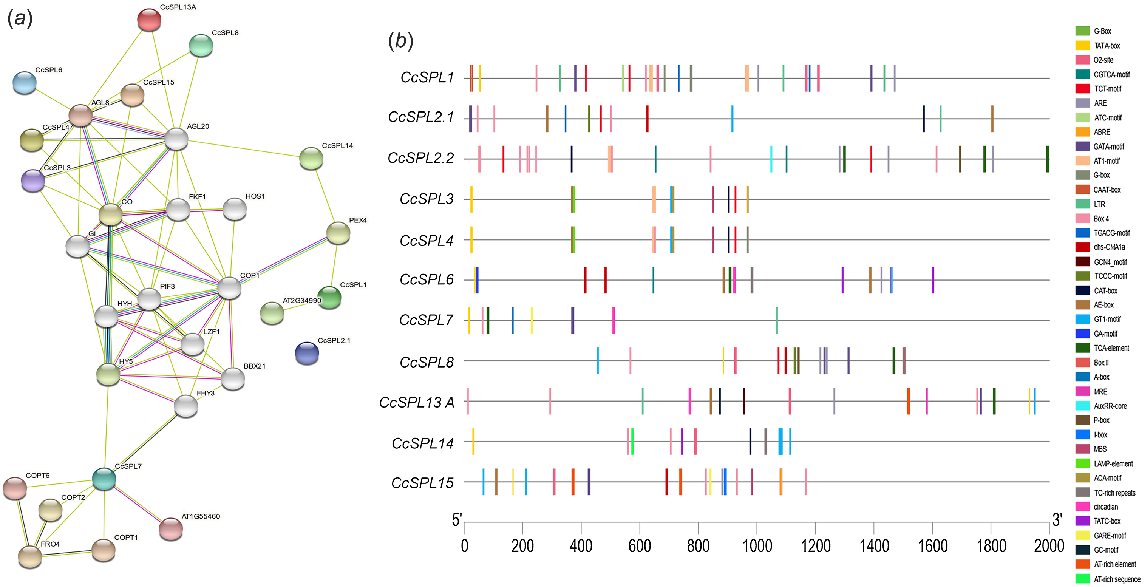
Cis-acting regulatory elements of CcSPL genes
Cis-acting regulatory elements are the main molecular switches for transcriptional regulatory activities of genes that affect a wide range of biological processes, i.e. hormone response, development and abiotic stress response (Yamaguchi-Shinozaki and Shinozaki 2005). Twenty-three types of hormones and stress-related cis-acting regulatory elements were predicted in promoter sequences analysis of CcSPL genes (Fig. 4b) (Supplementary file 1). Elements include ABRE (Abscisic Acid-Responsive Element), ARE (cis-acting regulatory element of anaerobic induction), BOX-4 (potentially involved in light responsiveness), CGTCA-MOTIF (which was involved in the hormonal MeJA-responsiveness), G-BOX (a trans-acting regulatory element with functions in light responsiveness), LTR (involved in low-temperature-responsive functions), 02-site (a trans-acting regulatory element involved in zein metabolism regulation), TGACG-motif (a trans-acting regulatory element involved in the MeJA-responsiveness), GCN4-motif (a trans-acting regulatory element related to endosperm expression), CAT-Box (a trans-acting regulatory element) and CAAT-box (cis-acting regulatory element related to meristem expression). All eleven members of the SPL gene family had the AT1 motif, GATA motif, TCT motif, AE box, GA motif, GT1 motif, TCCC motif, chs-CMA1a, LAMP element, box I and box II; each of these elements was a component of a light-responsive element (Mongkolsiriwatana et al. 2009).
Gene expression of CcSPLs in salt-tolerant and salt-susceptible pigeon pea genotypes under control and salt-treated roots
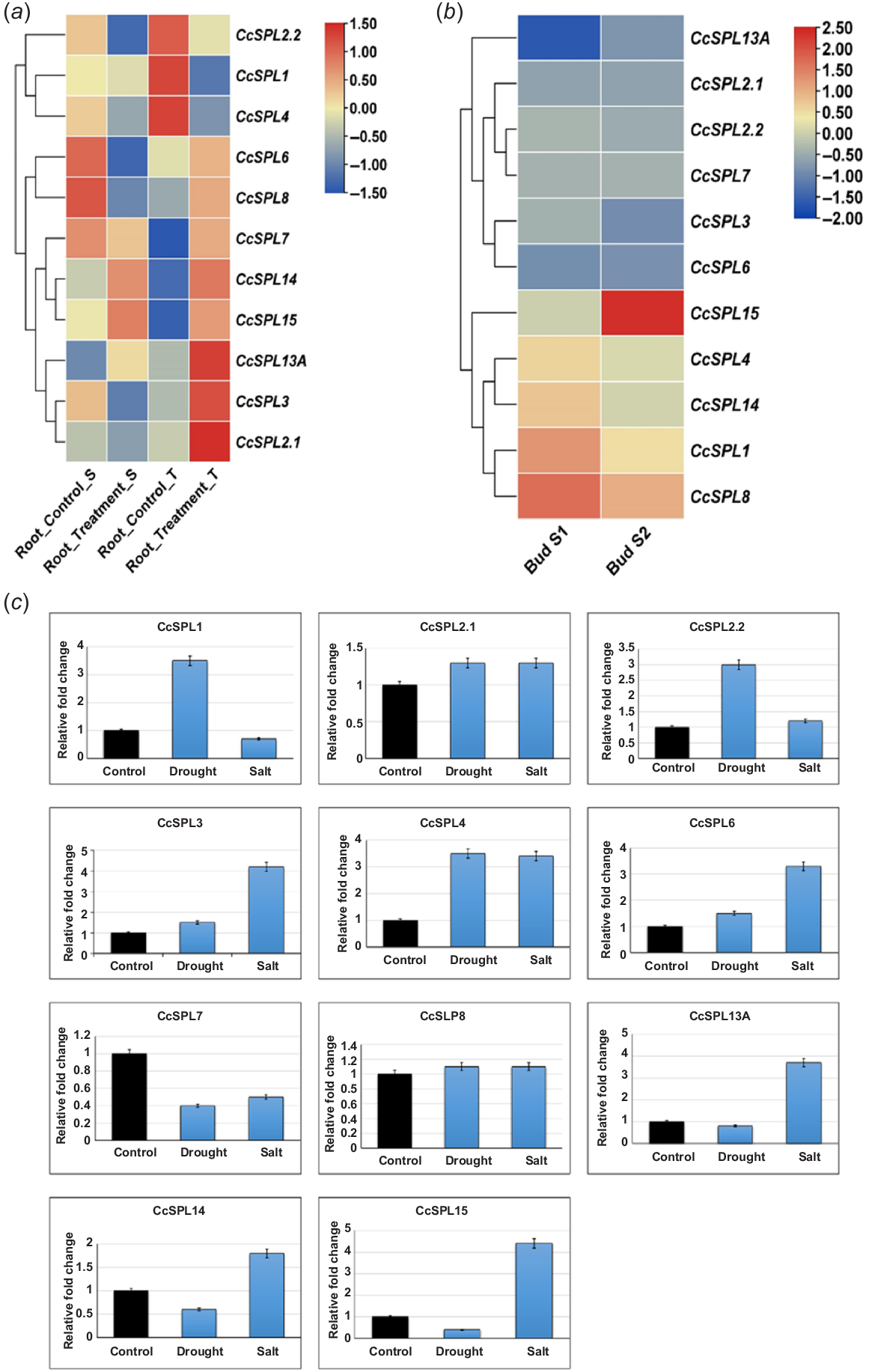
To gain insight into the functional properties of CcSPL family members, an RNA-seq analysis was performed using data from the NCBI SRA (Sequence Read Archive) database. A total of four runs were observed under bio project PRJNA382795. The current study involves the analysis of gene expression patterns of C. cajan root in two cultivars; ICP 7 (salt-tolerant pigeon pea genotype) and ICP 1071 (salt-susceptible pigeon pea genotype). A total of 39.25 gigabytes of experiment data was uploaded to SRA (SRP104063). The experimental data were used to evaluate the PKRM values using the Galaxy server-based tool and produced almost 61.04 gigabytes of data with more than 30 301 923 spots. The PKRM values were collected for all the CcSPL genes and the heatmap was generated using TB tools (Chen et al. 2020). In the heatmap, rows represent gene names and columns represent experiment values. The blue colour represents the downregulated genes, yellow represents no change of expression and red represents an upregulation (Fig. 5a). In the susceptible cultivar, CcSPL6 and CcSPL8 were downregulated whereas, CcSPL14 and CcSPL15 were upregulated under salt stress conditions. Similarly, in the salt-tolerant cultivar, all the CcSPLs were significantly upregulated in response to salt stress, except for CcSPL1, CcSPL2.2 and CcSPL4 (which were downregulated). It is important to notice that CcSPL14 and CcSPL15 were upregulated under salinity stress for both ICP 7 and ICP 1071 cultivars (Fig. 5a). In another SRA experiment, transcriptomes of two phases of unopened flowering buds of an isogenic C. cajan line ICPB 2043 were analysed. A total of eight runs were observed under bio project PRJNA634080. The expression of CcSPL genes can be divided into two distinct groups as highly expressed and weakly detected. The CcSPL1 and CcSPL8 were the most expressed transcripts at Bud Stage 1. Similarly, CcSPL8 and CcSPL15 were highly expressed at Bud Stage 2 as compared to other members of this family (Supplementary file 2) (Fig. 5b).
Gene expression analysis. (a) RNA-seq based expression profiles of CcSPL genes in response to salt stress in salt-susceptible and tolerant cultivars. (b) RNA-seq based expression profiles of CcSPL genes during two stages flower bud formation. The expression values [log2(FPKM)] are shown by the coloured bar. Red denotes high expression, whereas yellow denotes low expression; blue indicates no expression. Rectangles of various colours are used to denote the various CcSPL gene groupings. Table S1 lists the FPKM values of the CcSPL genes in various tissues. (c) SPL gene family relative qPCR test in response to abiotic (salt and drought) stresses. To produce an objective average value, the experiment was triplicated. In untreated plants, each gene had a default expression value of 1. Bars have been placed on each column to illustrate the standard error.
Gene expression analysis of CcSPL genes
To validate the expression of CcSPLs, real-time PCR-based expression quantification was performed for CcSPL genes in response to salt and drought stress (Hanafy et al. 2013). In response to drought stress, CcSPL1, 2.2 and 4 were significantly upregulated. On the other hand, CcSPL7, 14 and 15 were downregulated. The expression of other CcSPLs genes were not significantly modulated under drought stress conditions (Fig. 5c). In response to salt stress, CcSPL3, CcSPL4, CcSPL6, CcSPL13A and CcSPL15 were upregulated whereas, the expression of CcSPL7 was downregulated by salinity. It was observed that CcSPL4 is upregulated in response to both salt and drought stress conditions. When compared to control, CcSPL7 was shown to be downregulated under salt and drought stresses (Fig. 5c).
3D protein structure prediction and docking analysis
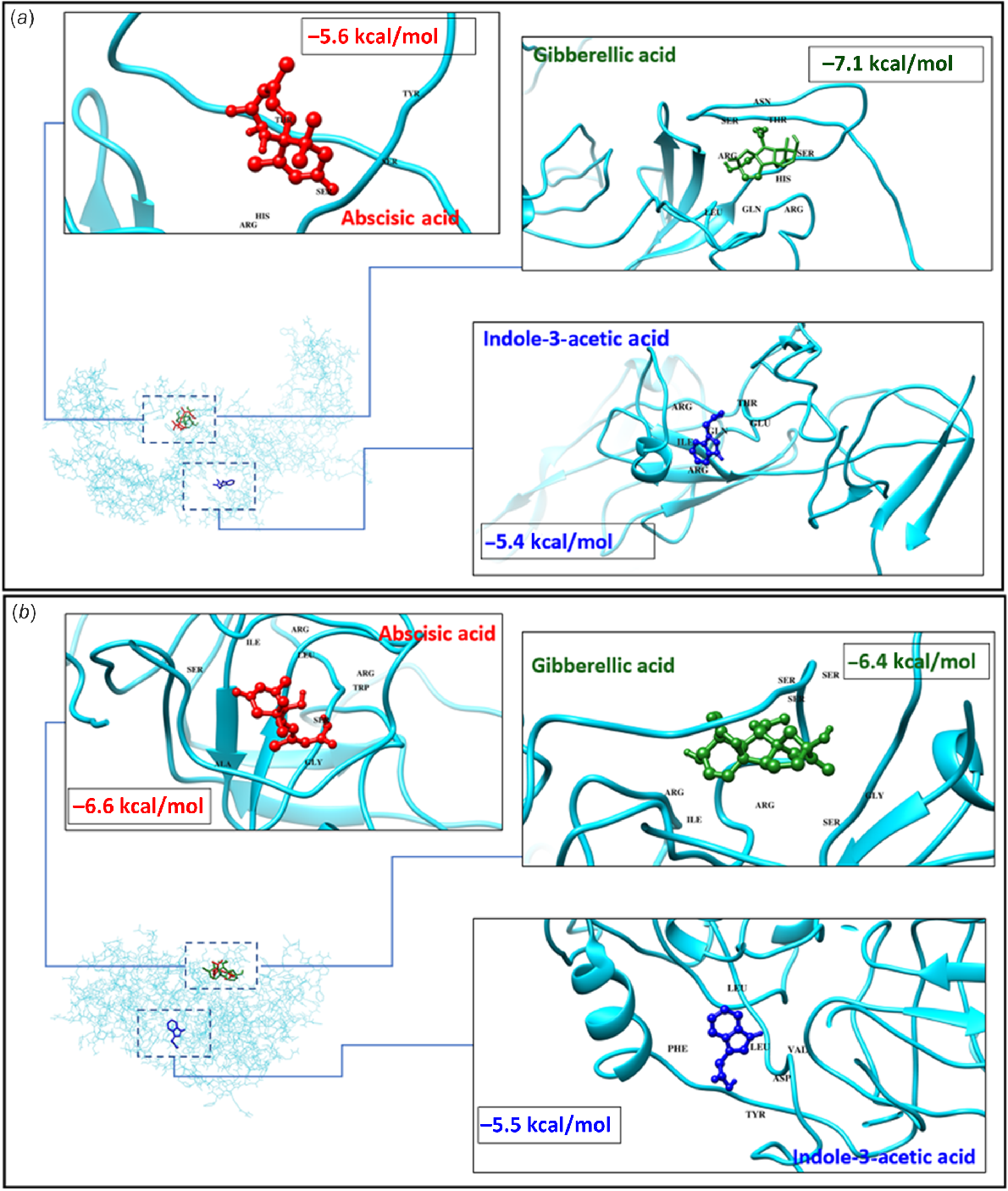
Protein 3D structures for all CcSPL proteins were predicted through I-TASER and the energy minimisation step was performed using Galaxy Mod-refiner. Since CcSPL3 and CcSPL14 protein expression were significantly modulated in response to environmental stimuli (Fig. 5), these were selected for structure prediction and docking analysis. Moreover, ABA and GA3 are important phytohormones for plant growth and stress response. Therefore, these two were selected as ligands in docking analysis. After the docking analysis, results were also collected for other CcSPL proteins and presented in the supplementary data (Supplementary files 3 and 4). The Fig. 6 displayed the 3D structure of CcSPL3 (Fig. 6a) and CcSPL14 (Fig. 6b) as well as the ligands attached to them. In Fig. 6a it can also be observed that ABA was attached to CcSPL3 at two Arg sites, two Glu sites, and at an Asn site with binding affinity of −4.4 kcal/mol, GA3 was attached at Arg, three Gln sites, Phe, Glu, Ile, Lys and Val sites with a binding affinity of −5.5 kcal/mol, and IAA was attached at Asp, Lys, Val, Glu and Arg with a binding affinity of −4.5 kcal/mol. Similarly, Fig. 6b showed ABA was attached to CcSPL14 at Ser, Ile, Arg, Leu, Trg, Ser and Gly sites with a binding affinity of −6.6 kcal/mol, GA3 was attached at two Arg sites, Ile, and multiple Ser sites with a binding affinity of −6.4 kcal/mol and IAA was attached at Phe, two Leu sites, Val, Tyr and Asp sites with a binding affinity of −5.5 kcal/mol.
Discussion
The SPL gene family plays a crucial role in a plant’s adaptation to abiotic stress and in its development (Preston and Hileman 2013). The functional characterisation of SPL genes has been performed in only a limited number of model and economicaly important crop plants (Chao et al. 2017, 2019; Zhong et al. 2019; Xu et al. 2020). However, SPLs in C. cajan were yet to be explored. The current study presents the first comprehensive report of the SPL gene family in neglected legumes. Therefore, it can be used as a reference to explore SPLs in related legumes for exploring novel avenues to improve crop plants. A total of 11 SPL genes were identified in C. cajan as compared to 17 SPLs in A. thaliana. Similarly, 9, 41, 11, 12 and 11 SPL genes were reported in C. arietinum, G. max, P. vulgaris, V. unguiculata and Arachis hypogaea, respectively. The genome sizes of Arabidopsis, C. cajan, C. arietinum, G. max, P. vulgaris, V. unguiculata and A. hypogaea are 119.75 Mb, 592.971 Mb, 511.684 Mb, 993.002 Mb, 452.733 Mb, 519.067 and 2529.19 Mb, respectively. The number of SPL genes are not correlated to genome size. It is already reported that SPL genes may have evolved in a species-specific manner (Yu et al. 2021).
According to phylogenetic analysis, the SPL family TFs are clustered into nine groups. These findings are in accordance with previous studies (Zhang et al. 2015; Mao et al. 2016; Yu et al. 2021). The CcSPL1 protein possesses an ANK-repeats domain (no other CcSPL possesses the ANK domain). The ANK domain is also present in 3 SPLs in Betula luminifera SPLs in Gossypium hirsutum, and six SPLs in Zea mays (Mao et al. 2016; Li et al. 2018; Ahmed et al. 2023). Moreover, the G1 members possess the ANK-repeat domain (Fig. 1). The ANK-repeat domain promotes protein–protein interaction (Fig. 4). When the conserved domains of the proteins were identified and compared it was revealed that CcSPL proteins contained dual zinc-binding domains Zn1, Zn2 (CCCH and CCHC) and one NLS (KR) site. Besides, this classification was generally dependent on gene structure motif analysis, and localisations as described in Figs 3 and 4. In the present study, 23 types of hormones and stress-related cis-acting regulatory elements were identified in 2000 bp upstream sequences of CcSPL genes, which include the promoter and UTR region (Brogan et al. 1999; Rehman et al. 2022). Phytohormone ABA is involved in stress response and gibberellins are involved in flowering (Abeed and Dawood 2020). Therefore, the presence of respective transcription factor binding sites suggests the involvement of CcSPLs in phytohormone-mediated plant responses.
To further investigate the likely function of the CcSPL gene family in C. cajan response to salt stress and the development of bud, the expression patterns of the 11 CcSPL members in five different tissues were examined based on previously published transcriptome data (Fig. 5). Drought stress-responsive CcSPL1, 2.2 and 4 are homologues of AtSPL1, AtSPL2 and AtSPL4 respectively (Fig. 1). The AtSPL1 and AtSPL12 act redundantly in thermotolerance (Chao et al. 2017). Similarly, AtSPL1 and AtSPL2 are involved in heat-shock memory (Stief et al. 2014). The AtSPL4 is involved in aging and photoperiodic signal transduction (Jung et al. 2016). Similarly, homologues of salt stress-responsive CcSPL3, CcSPL4, CcSPL6, CcSPL13A and CcSPL15 are involved in environmental stress responses (Stief et al. 2014; Jung et al. 2016; Yu et al. 2021). These reports strengthen the stress-responsive role of these SPL gene family members. Docking was performed on two highly expressed genes, one of which was CcSPL3 and the other was CcSPL14. Successful docking of ABA, GA3 and IAA with CcSPL3 and CcSPL14 predicts a significant impact on these hormones. The overexpression of OsmiR156k reduced rice tolerance to cold stress and it also downregulated SPL3, SPL14 and SPL17 (Cui et al. 2015). Moreover, SPL14 overexpression accelerates the flowering of Aechmea fasciatai (Lei et al. 2018). Two microRNAs (miRNA156 and miRNA529) are important regulators of SPLs in plants (Ling and Zhang 2012). Therefore, future studies are expected to explore the relationship between miRNAs and the stress response of SPLs in C. cajan.
Conclusion
In the current study, 11 SPL genes were identified in C. cajan. The phylogenetic analysis classified CcSPLs into nine groups. Further, the identification of gene structure, motif analysis, domain analysis, and presence of cis-regulatory elements in the SPL family members were studied. The RNA-Seq data analysis revealed differential regulation of CcSPL2.1, 3, 13 A, 14 and 15 in response to salt stress. Real-time qPCR validated the transcriptional response of these genes to drought and salt stress. Molecular docking was performed against the proteins of the two highly expressed genes (CcSPL3 and CcSPL14) with the three ligands abscisic acid (ABA), gibberellic acid (GA3) and indole-3-acetic acid (IAA). Afterward, their binding affinity was obtained and the 3D structures were predicted. ABA and IAA are important phytohormones, therefore, the findings of the current study will serve as a basis for further functional insights into the SPL gene family.
Data availability
The data presented in the study are included in the article and in supplementary material.
Conflicts of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflicts of interest.
Declaration of funding
No specific funding has been received to conduct or publish present research findings.
Author contributions
FA designed the research; TS, AR, MW and FA executed the experiment; MQ, FA, MA and MFA helped in data analysis and original manuscript write-up. FA supervised the research experiment and review. KA, AMA, NN and SE provided technical expertise to strengthen the basic idea of the research. NN, AMA, MFA, SE and AR helped in literature review, data analysis, critical review of the manuscript and funding acquisition for this publication. TS, FA, AHAA, NN, AA and SF helped in revision and streamlining the article as per reviewers comments. All authors helped with proofreading in the revision of the article.
Acknowledgements
The authors extend their appreciation to the Researchers Supporting Project number (RSP-2024R369), King Saud University, Riyadh, Saudi Arabia.
References
Abeed AHA, Dawood MFA (2020) Comparative impact of different iso-osmotic solutions on osmotic adjustment in Gossypium barbadense. Global NEST Journal 22, 75-84.
| Crossref | Google Scholar |
Abeed AHA, Ali M, Ali EF, Majrashi A, Eissa MA (2021) Induction of Catharanthus roseus secondary metabolites when Calotropis procera was used as bio-stimulant. Plants 10, 1623.
| Crossref | Google Scholar | PubMed |
Ahmed S, Rashid MAR, Zafar SA, Azhar MT, Waqas M, Uzair M, et al. (2021) Genome-wide investigation and expression analysis of APETALA-2 transcription factor subfamily reveals its evolution, expansion and regulatory role in abiotic stress responses in Indica Rice (Oryza sativa L. ssp. indica). Genomics 113, 1029-1043.
| Crossref | Google Scholar | PubMed |
Ahmed T, Masood HA, Noman M, AL-Huqail AA, Alghanem SMS, Khan MM, Muhammad S, Manzoor N, Rizwan M, Qi X, Abeed AHA, Li B (2023) Biogenic silicon nanoparticles mitigate cadmium (Cd) toxicity in rapeseed (Brassica napus L.) by modulating the cellular oxidative stress metabolism and reducing Cd translocation. Journal of Hazardous Materials 459, 132070.
| Crossref | Google Scholar | PubMed |
Ali MA, Azeem F, Nawaz MA, Acet T, Abbas A, Imran QM, et al. (2018) Transcription factors WRKY11 and WRKY17 are involved in abiotic stress responses in Arabidopsis. Journal of Plant Physiology 226, 12-21.
| Crossref | Google Scholar | PubMed |
Bailey TL, Johnson J, Grant CE, Noble WS (2015) The MEME Suite. Nucleic Acids Research 43, W39-W49.
| Crossref | Google Scholar | PubMed |
Brogan IJ, Khan N, Isaac K, Hutchinson JA, Pravica V, Hutchinson IV (1999) Novel polymorphisms in the promoter and 5′ UTR regions of the human vascular endothelial growth factor gene. Human Immunology 60, 1245-1249.
| Crossref | Google Scholar | PubMed |
Cao R, Guo L, Ma M, Zhang W, Liu X, Zhao H (2019) Identification and functional characterization of Squamosa promoter binding protein-like gene TaSPL16 in wheat (Triticum aestivum L.). Frontiers in Plant Science 10, 212.
| Crossref | Google Scholar |
Cardon G, Höhmann S, Klein J, Nettesheim K, Saedler H, Huijser P (1999) Molecular characterisation of the Arabidopsis SBP-box genes. Gene 237, 91-104.
| Crossref | Google Scholar | PubMed |
Chao L-M, Liu Y-Q, Chen D-Y, Xue X-Y, Mao Y-B, Chen X-Y (2017) Arabidopsis transcription factors SPL1 and SPL12 confer plant thermotolerance at reproductive stage. Molecular Plant 10, 735-748.
| Crossref | Google Scholar | PubMed |
Chen C, Chen H, Zhang Y, Thomas HR, Frank MH, He Y, et al. (2020) TBtools: an integrative toolkit developed for interactive analyses of big biological data. Molecular Plant 13, 1194-1202.
| Crossref | Google Scholar | PubMed |
Crooks GE, Hon G, Chandonia J-M, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Research 14, 1188-1190.
| Crossref | Google Scholar | PubMed |
Cui N, Sun X, Sun M, Jia B, Duanmu H, Lv D, et al. (2015) Overexpression of OsmiR156k leads to reduced tolerance to cold stress in rice (Oryza Sativa). Molecular Breeding 35, 214.
| Crossref | Google Scholar |
Dar WD (2009) ICRISAT surging ahead with innovations: a compendium of speeches and presentations. ICRISAT, International Crops Research Institute for the Semi-Arid Tropics. Available at https://oar.icrisat.org/1358/
Goddard TD, Huang CC, Ferrin TE (2005) Software extensions to UCSF chimera for interactive visualization of large molecular assemblies. Structure 13, 473-482.
| Crossref | Google Scholar | PubMed |
Halim SA, Waqas M, Khan A, Al-Harrasi A (2021) In silico prediction of novel inhibitors of SARS-CoV-2 main protease through structure-based virtual screening and molecular dynamic simulation. Pharmaceuticals 14, 896.
| Crossref | Google Scholar | PubMed |
Hanafy MS, El-Banna A, Schumacher HM, Jacobsen H-J, Hassan FS (2013) Enhanced tolerance to drought and salt stresses in transgenic faba bean (Vicia faba L.) plants by heterologous expression of the PR10a gene from potato. Plant Cell Reports 32, 663-674.
| Crossref | Google Scholar | PubMed |
Hou H, Li J, Gao M, Singer SD, Wang H, Mao L, et al. (2013) Genomic organization, phylogenetic comparison and differential expression of the SBP-box family genes in grape. PLoS ONE 8, e59358.
| Crossref | Google Scholar | PubMed |
Hu B, Jin J, Guo A-Y, Zhang H, Luo J, Gao G (2015) GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics 31, 1296-1297.
| Crossref | Google Scholar | PubMed |
Ioannidi E, Rigas S, Tsitsekian D, Daras G, Alatzas A, Makris A, et al. (2016) Trichome patterning control involves TTG1 interaction with SPL transcription factors. Plant Molecular Biology 92, 675-687.
| Crossref | Google Scholar | PubMed |
Jorgensen SA, Preston JC (2014) Differential SPL gene expression patterns reveal candidate genes underlying flowering time and architectural differences in Mimulus and Arabidopsis. Molecular Phylogenetics and Evolution 73, 129-139.
| Crossref | Google Scholar | PubMed |
Jung J-H, Lee H-J, Ryu JY, Park C-M (2016) SPL3/4/5 integrate developmental aging and photoperiodic signals into the FT-FD module in Arabidopsis flowering. Molecular Plant 9, 1647-1659.
| Crossref | Google Scholar | PubMed |
Kim S, Chen J, Cheng T, Gindulyte A, He J, He S, et al. (2019) PubChem 2019 update: improved access to chemical data. Nucleic Acids Research 47, D1102-D1109.
| Crossref | Google Scholar | PubMed |
Kropat J, Tottey S, Birkenbihl RP, Depège N, Huijser P, Merchant S (2005) A regulator of nutritional copper signaling in Chlamydomonas is an SBP domain protein that recognizes the GTAC core of copper response element. Proceedings of the National Academy of Sciences 102, 18730-18735.
| Crossref | Google Scholar |
Lei M, Li Z-Y, Wang J-B, Fu Y-L, Ao M-F, Xu L (2018) Constitutive expression of Aechmea fasciata SPL14 (AfSPL14) accelerates flowering and changes the plant architecture in Arabidopsis. International Journal of Molecular Sciences 19, 2085.
| Crossref | Google Scholar | PubMed |
Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, et al. (2002) PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Research 30, 325-327.
| Crossref | Google Scholar | PubMed |
Li X-Y, Lin E-P, Huang H-H, Niu M-Y, Tong Z-K, Zhang J-H (2018) Molecular Characterization of SQUAMOSA PROMOTER BINDING PROTEIN-LIKE (SPL) Gene Family in Betula luminifera. Frontiers in Plant Science 9, 608.
| Crossref | Google Scholar |
Ling L-Z, Zhang S-D (2012) Exploring the evolutionary differences of SBP-box genes targeted by miR156 and miR529 in plants. Genetica 140, 317-324.
| Crossref | Google Scholar | PubMed |
Liu M, Sun W, Ma Z, Huang L, Wu Q, Tang Z, et al. (2019) Genome-wide identification of the SPL gene family in Tartary Buckwheat (Fagopyrum tataricum) and expression analysis during fruit development stages. BMC Plant Biology 19, 299.
| Crossref | Google Scholar |
Mao H-D, Yu L-J, Li Z-J, Yan Y, Han R, Liu H, et al. (2016) Genome-wide analysis of the SPL family transcription factors and their responses to abiotic stresses in maize. Plant Gene 6, 1-12.
| Crossref | Google Scholar |
Mongkolsiriwatana C, Pongtongkam P, Peyachoknagul S (2009) In silico promoter analysis of photoperiod-responsive genes identified by DNA microarray in rice (Oryza sativa L.). Agriculture and Natural Resources 43, 164-177.
| Google Scholar |
Nadeem M, Khan AA, Nadeem J, Khan AA, Fatima U (2023) Cloning and characterization of Trichoderma glucanase gene for plant transformation. International Journal of Agriculture and Biosciences 12(1), 31-46.
| Crossref | Google Scholar |
Nguyen L-T, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Molecular Biology and Evolution 32, 268-274.
| Crossref | Google Scholar | PubMed |
Noman MU, Azhar S (2023) Metabolomics, a potential way to improve abiotic stresses tolerance in cereal crops. International Journal of Agriculture and Biosciences 12(1), 47-55.
| Crossref | Google Scholar |
Palmeri D, Malim MH (1999) Importin β can mediate the nuclear import of an arginine-rich nuclear localization signal in the absence of importin α. Molecular and Cellular Biology 19, 1218-1225.
| Crossref | Google Scholar | PubMed |
Pasquier J, Cabau C, Nguyen T, Jouanno E, Severac D, Braasch I, et al. (2016) Gene evolution and gene expression after whole genome duplication in fish: the PhyloFish database. BMC Genomics 17, 368.
| Crossref | Google Scholar |
Preston JC, Hileman LC (2013) Functional evolution in the plant SQUAMOSA-PROMOTER BINDING PROTEIN-LIKE (SPL) gene family. Frontiers in Plant Science 4, 80.
| Crossref | Google Scholar |
Rehman I, Shaukat F, Anwar Z, Ijaz A, Nadeem R (2022) Applications of Crispr/Cas system in plants. International Journal of Agriculture and Biosciences 11(4), 231-237.
| Crossref | Google Scholar |
Sherry ST, Ward M-H, Kholodov M, Baker J, Phan L, Smigielski EM, et al. (2001) dbSNP: the NCBI database of genetic variation. Nucleic Acids Research 29, 308-311.
| Crossref | Google Scholar | PubMed |
Stief A, Altmann S, Hoffmann K, Pant BD, Scheible W-R, Bäurle I (2014) Arabidopsis miR156 regulates tolerance to recurring environmental stress through SPL transcription factors. The Plant Cell 26, 1792-1807.
| Crossref | Google Scholar | PubMed |
Sun H, Mei J, Zhao W, Hou W, Zhang Y, Xu T, et al. (2022) Phylogenetic analysis of the SQUAMOSA promoter-binding protein-like genes in four Ipomoea species and expression profiling of the IbSPLs during storage root development in sweet potato (Ipomoea batatas). Frontiers in Plant Science 12, 801061.
| Crossref | Google Scholar |
Swarbreck D, Wilks C, Lamesch P, Berardini TZ, Garcia-Hernandez M, Foerster H, et al. (2007) The Arabidopsis Information Resource (TAIR): gene structure and function annotation. Nucleic Acids Research 36, D1009-D1014.
| Crossref | Google Scholar | PubMed |
Takahashi H, Fujishima T, Koba H, Murakami S, Kurokawa K, Shibuya Y, et al. (2000) Serum surfactant proteins A and D as prognostic factors in idiopathic pulmonary fibrosis and their relationship to disease extent. American Journal of Respiratory and Critical Care Medicine 162, 1109-1114.
| Crossref | Google Scholar | PubMed |
The UniProt Consortium (2015) UniProt: a hub for protein information. Nucleic Acids Research 43, D204-D212.
| Crossref | Google Scholar |
Varshney RK, Chen W, Li Y, Bharti AK, Saxena RK, Schlueter JA, et al. (2012) Draft genome sequence of pigeonpea (Cajanus cajan), an orphan legume crop of resource-poor farmers. Nature Biotechnology 30, 83-89.
| Crossref | Google Scholar |
Waqas M, Azhar MT, Rana IA, Azeem F, Ali MA, Nawaz MA, et al. (2019) Genome-wide identification and expression analyses of WRKY transcription factor family members from chickpea (Cicer arietinum L.) reveal their role in abiotic stress-responses. Genes & Genomics 41, 467-481.
| Crossref | Google Scholar | PubMed |
Waqas M, Haider A, Rehman A, Qasim M, Umar A, Sufyan M, et al. (2021) Immunoinformatics and molecular docking studies predicted potential multiepitope-based peptide vaccine and novel compounds against novel SARS-CoV-2 through virtual screening. BioMed Research International 2021, 1596834.
| Crossref | Google Scholar |
Wu G, Poethig RS (2006) Temporal regulation of shoot development in Arabidopsis thaliana by miR156 and its target SPL3. Development 133, 3539-3547.
| Crossref | Google Scholar | PubMed |
Wu G, Park MY, Conway SR, Wang J-W, Weigel D, Poethig RS (2009) The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis. Cell 138, 750-759.
| Crossref | Google Scholar | PubMed |
Xu Y, Xu H, Wall MM, Yang J (2020) Roles of transcription factor SQUAMOSA promoter binding protein-like gene family in papaya (Carica papaya) development and ripening. Genomics 112, 2734-2747.
| Crossref | Google Scholar | PubMed |
Xue X-Y, Zhao B, Chao L-M, Chen D-Y, Cui W-R, Mao Y-B, et al. (2014) Interaction between two timing microRNAs controls trichome distribution in Arabidopsis. PLoS Genetics 10, e1004266.
| Crossref | Google Scholar | PubMed |
Yamaguchi-Shinozaki K, Shinozaki K (2005) Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends in Plant Science 10, 88-94.
| Crossref | Google Scholar | PubMed |
Yang Z, Wang X, Gu S, Hu Z, Xu H, Xu C (2008) Comparative study of SBP-box gene family in Arabidopsis and rice. Gene 407, 1-11.
| Crossref | Google Scholar | PubMed |
Yu N, Cai W-J, Wang S, Shan C-M, Wang L-J, Chen X-Y (2010) Temporal control of trichome distribution by microRNA156-targeted SPL genes in Arabidopsis thaliana. The Plant Cell 22, 2322-2335.
| Crossref | Google Scholar | PubMed |
Yu P, Shinde H, Dudhate A, Tsugama D, Gupta SK, Liu S, et al. (2021) Genome-wide investigation of SQUAMOSA promoter binding protein-like transcription factor family in pearl millet (Pennisetum glaucum (L) R. Br.). Plant Gene 27, 100313.
| Crossref | Google Scholar |
Zhang X, Dou L, Pang C, Song M, Wei H, Fan S, et al. (2015) Genomic organization, differential expression, and functional analysis of the SPL gene family in Gossypium hirsutum. Molecular Genetics and Genomics 290, 115-126.
| Crossref | Google Scholar | PubMed |
Zhong H, Kong W, Gong Z, Fang X, Deng X, Liu C, et al. (2019) Evolutionary analyses reveal diverged patterns of SQUAMOSA promoter binding protein-like (SPL) gene family in Oryza genus. Frontiers in Plant Science 10, 565.
| Crossref | Google Scholar |